The Dynamic Transcriptome: Using Clustered Time Points to Tease Apart Rice Tiller Angle Control
Given the importance of branch angle in determining overall plant form, we know surprisingly little about how those angles are controlled. Branch angle clearly has genetic underpinnings but also responds strongly to light and gravity, as well as to water availability and other stimuli (reviewed in Roychoudhry and Kepinski, 2015). Analysis of tiller angle in rice (Oryza sativa) led to the discovery of LAZY1 (LA1), orthologs of have since been shown to determine shoot branch angle in both monocots and dicots. Auxin is known to be a key player in this process. Beyond that, several quantitative trait loci have been linked to tiller angle, but the strong effects of field conditions have hindered efforts to use them for breeding.
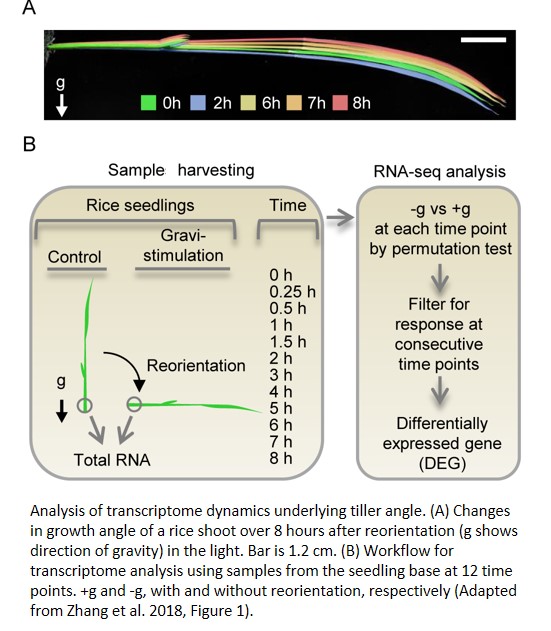
Now, Zhang et al. (2018) have developed a high-throughput approach to explore rice tiller angle, leading to the identification of both upstream and downstream pathway components. The authors took advantage of the likely connection between tiller angle and gravistimulation to generate a more controlled environment under which to find genetic factors influencing rice tiller angle. To this end, they rotated seedlings to impose gravistimulation in the light and characterized the transcriptome at the base of treated (and control) shoots at 12 time points during the next 8 hours, at which point upward growth could be seen in gravistimulated shoots (see Figure). Based on principal component analysis, the transcriptomes of the treated shoots became different from those of the control after 2 hours, prompting the authors to divide the response into early (up to 2 h) and late (3–8 h) stages.
 The use of many time points allowed the authors to limit their definition of differentially expressed genes (DEGs) to those that were different from the control at two consecutive time points. They found 98 DEGs in the early stage, 3246 in the late stage and 860 so-called dual DEGs that were altered in both stages. Within these groups, Zhang et al. formed clusters based on expression patterns to gain a sense of what processes might be involved in each stage. Further analysis allowed the authors to assess the likely relationships between clusters and build a predicted regulatory network, highlighting candidates for experimental validation.
The use of many time points allowed the authors to limit their definition of differentially expressed genes (DEGs) to those that were different from the control at two consecutive time points. They found 98 DEGs in the early stage, 3246 in the late stage and 860 so-called dual DEGs that were altered in both stages. Within these groups, Zhang et al. formed clusters based on expression patterns to gain a sense of what processes might be involved in each stage. Further analysis allowed the authors to assess the likely relationships between clusters and build a predicted regulatory network, highlighting candidates for experimental validation.
Consistent with the known role of auxin in gravitropism and branch angle control, auxin-related genes were enriched among late and dual DEGs. Zhang et al. looked within their dataset for genes that responded early and thus might feed into the auxin responses. This led them to HEAT STRESS TRANSCRIPTION FACTOR 2D (HSFA2D), and the authors found, indeed, that the hsfa2d mutant had increased tiller angle in the field as well as reduced gravitropic responses. The mutant also showed decreased lateral auxin transport under gravistimulation, indicating HSFA2D functions upstream of auxin.
Given that LA1 also functions upstream of auxin to mediate tiller angle and gravitropic responses, the authors examined the relationship between LA1 and HSFA2D, finding evidence that HSFA2D functions upstream of LA1 in the same pathway to induce lateral transport of auxin.
Having established these upstream players, Zhang et al. went on to mine their temporal transcriptome data for factors potentially acting downstream of auxin gradients in tiller angle control. They identified WOX6 and WOX11 as auxin-responsive transcription factor genes and provided evidence that they function redundantly downstream of LA1 and HSFA2D.
These findings highlight the effectiveness of using clustered time points to elucidate transcriptome dynamics, a strategy that could be applied to other traits that have been hard to dissect.
REFERENCES
Roychoudhry, S., and Kepinski, S. (2015). Shoot and root branch growth angle control-the wonderfulness of lateralness. Curr. Opin. Plant Biol. 23: 124-131.
Zhang, N., Yu, H., Yu, H. Cai, Y. Huang, L., Xu, C., Xiong, G., Meng, X., Wang, J., Chen, H., Liu, G., Jing, Y., Yuan, Y., Liang, Y., Li, S., Smith, S.M., Li, J., and Wang, Y. (2018). A Core Regulatory Pathway Controlling Rice Tiller Angle Mediated by the LAZY1-dependent Asymmetric Distribution of Auxin. Plant Cell. doi: 10.1105/tpc.18.00063.



