Sugar architect: the Brassicaceae pathogen Clubroot manipulates plants on multiple levels to secure sucrose supply
The soil-borne pathogen Plasmodiophora brassicae can infect most members of the Brassicaceae family. The infections, which can lead to extensive crop losses, typically involve development of galls in the underground tissues of the plant, giving the pathogen its common name, ‘clubroot’.
Although not a mandatory part of the pathogen cycle, these galls act as a protected hub for pathogen development and maturation, with formation influencing the number of spores released into the soil for further infection cycles (Malinowski et al., 2012).
Galls have long been known to be a major sink for plant-derived carbon, and gall formation has previously been linked to increased numbers of phloem cells. Furthermore, the phloem differentiation mutant cle41, has been shown to develop smaller galls (Malinowski et al., 2012). This proposed ‘critical link’ between pathogen performance and the ability for the pathogen to sequester carbon from its host was further investigated in a recent study by Walerowski and colleagues (2018).
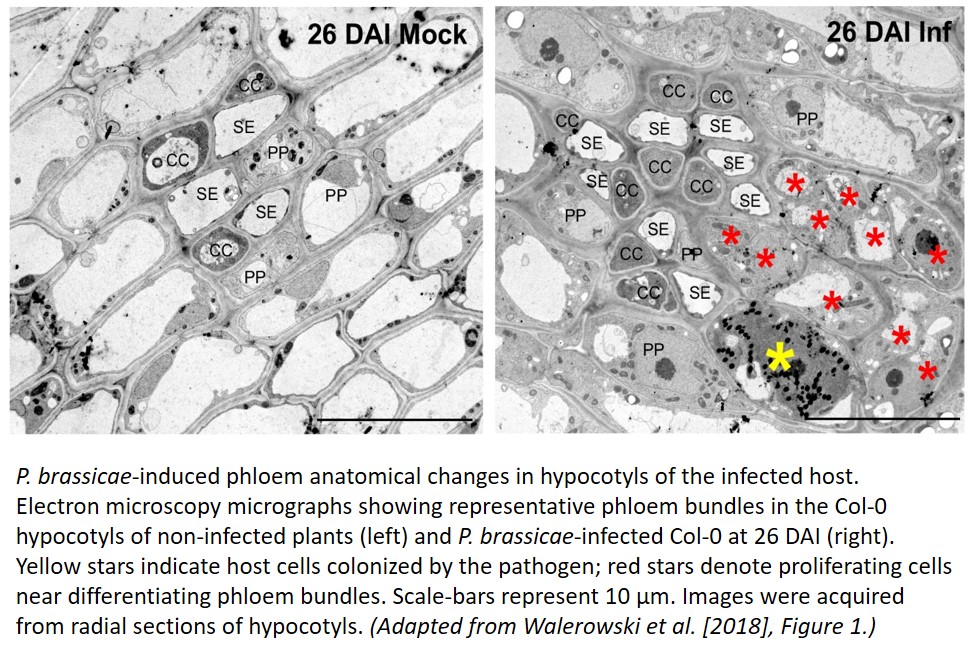
Following infection of Arabidopsis thaliana with P. brassicacae, the authors observed an increase in the number of all phloem cells with no marked changes in the ratios of cell types, indicating that reprogramming occurs early in the phloem differentiation process (see Figure). Promoter:GUS transgenic lines, used to investigate the cell-specific patterns of factors known to be involved early in phloem differentiation revealed phloem specific accumulation of OPS and CVP2 during the early proliferative stage of gall formation (16 days after infection). Interestingly, phloem development loss-of-function mutants ops-2, cvp2-1/cvl1-1 and brx-2 were not only unable to increase their phloem complexity to infected wild-type levels upon infection, as expected, but also showed disrupted gall anatomy. This resulted in a presumably compensative increase in the speed of the pathogenic life cycle, but also in premature death of the host. Together, these suggest that early reprogramming may not only benefit the pathogen, but may also allow the host to survive to the seed-formation stage.
In addition to the changes in phloem architecture, the authors observed an impact on enzymes involved in carbohydrate metabolism and transport. This included a transcriptional level decline in most invertases, and increases in sucrose synthases (such as SUS2 and SUS3) as well as the sucrose and hexose sugar transporters SWEET11 and SWEET12. The infection induced increases in both transporters could be supported by GUS-reporter assays.
Fourier transform infrared microspectroscopy, used to visualize the carbohydrate distribution within tissues showed the formation of a strong carbon sink in the central xylem region and ring of cambial cells, in infected wild-type plants with this response occurring at reduced magnitude in the sweet mutants. This in turn resulted in significantly lower accumulation of sucrose in the region. In line with this finding, sweet11:12 double mutants showed delays in pathogen development. Nonetheless, the host plants showed the same increases in phloem complexity associated with infection as displayed by the wild-type, and no changes in plant mortality.
These findings highlight that both levels of reprogramming– transporters and phloem architecture– are crucial for the successful progression of the disease, but that early remodeling of the phloem may also be an important part of host survival, potentially reflecting the diverse range of substances transported in these vessels.
References
Malinowski, R., Smith, J. A., Fleming, A. J., Scholes, J. D. & Rolfe, S. A. 2012. Gall formation in clubroot-infected Arabidopsis results from an increase in existing meristematic activities of the host but is not essential for the completion of the pathogen life cycle. Plant Journal, 71, 226-238.
Walerowski, P., Gündel, A., Yahaya, N., Truman, W., Sobczak, M., Olszak, M., Rolfe, S., Borisjuk, L. & Malinowski, R. 2018. Clubroot Disease Stimulates Early Steps of Phloem Differentiation and Recruits SWEET Sucrose Transporters within Developing Galls. Plant Cell. 10.1105/tpc.18.00283.