Direct and indirect visualization of bacterial effector delivery into diverse plant cell types during infection
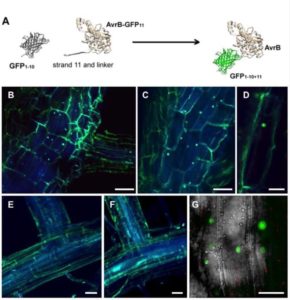 Bacterial effectors are proteins produced by bacteria and introduced into their hosts, where the effectors support successful infection by the pathogen. Effectors can function in diverse cells and cell compartments, but many studies of effector localization have relied on overexpression systems which may have artifacts. Henry et al. used the GFP strand system to visualize effector delivery during natural infection. In this system, plants are stably transformed with a gene encoding most but not all of the GFP protein, and the infecting bacteria express their effectors fused to the missing strand of the GFP structure. The GFP self-assembles in vivo, leading to fluorescence where the effector is delivered. This robust system revealed distinct effector localizations for various effectors and bacteria. Plant Cell 10.1105/tpc.17.0002 See also Park et al.
Bacterial effectors are proteins produced by bacteria and introduced into their hosts, where the effectors support successful infection by the pathogen. Effectors can function in diverse cells and cell compartments, but many studies of effector localization have relied on overexpression systems which may have artifacts. Henry et al. used the GFP strand system to visualize effector delivery during natural infection. In this system, plants are stably transformed with a gene encoding most but not all of the GFP protein, and the infecting bacteria express their effectors fused to the missing strand of the GFP structure. The GFP self-assembles in vivo, leading to fluorescence where the effector is delivered. This robust system revealed distinct effector localizations for various effectors and bacteria. Plant Cell 10.1105/tpc.17.0002 See also Park et al.










Leave a Reply
Want to join the discussion?Feel free to contribute!