Replaying the evolutionary tape to investigate subgenome dominance in allopolyploid Brassica napus (bioRxiv)
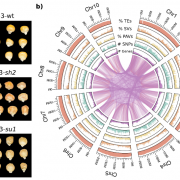
 Following interspecific hybridization, one of the two parental genomes (aka subgenomes) tends to become dominant (more highly retained and expressed). What determines which of the subgenomes will become dominant, or is it random? To explore this question, Bird et al. made several crosses between Brassica rapa (A genome) and Brassica oleracea (C genome), recapitulating the several-thousand year old hybridization event that led to Brassica napus, an allopolyploid hybrid (AACC). The authors examined six resynthesized polyploid lines across ten generations with DNA-seq, RNA-seq, and Bisulfite-seq data to characterize gene dosage, expression, and methylation differences. Across independent lines and generations, the authors found a bias towards C subgenome dominance. They conclude that, ” ‘replaying the evolutionary tape’ in allopolyploids results in repeatable and predictable subgenome expression dominance patterns based on preexisting genetic differences among the parental species.” (Summary by Mary Williams) bioRxiv
Following interspecific hybridization, one of the two parental genomes (aka subgenomes) tends to become dominant (more highly retained and expressed). What determines which of the subgenomes will become dominant, or is it random? To explore this question, Bird et al. made several crosses between Brassica rapa (A genome) and Brassica oleracea (C genome), recapitulating the several-thousand year old hybridization event that led to Brassica napus, an allopolyploid hybrid (AACC). The authors examined six resynthesized polyploid lines across ten generations with DNA-seq, RNA-seq, and Bisulfite-seq data to characterize gene dosage, expression, and methylation differences. Across independent lines and generations, the authors found a bias towards C subgenome dominance. They conclude that, ” ‘replaying the evolutionary tape’ in allopolyploids results in repeatable and predictable subgenome expression dominance patterns based on preexisting genetic differences among the parental species.” (Summary by Mary Williams) bioRxiv