Plants balance nucleotide availability with ribosome production
By Michael Busche, University of California-Berkeley, and Jacob O. Brunkard, University of Wisconsin-Madison
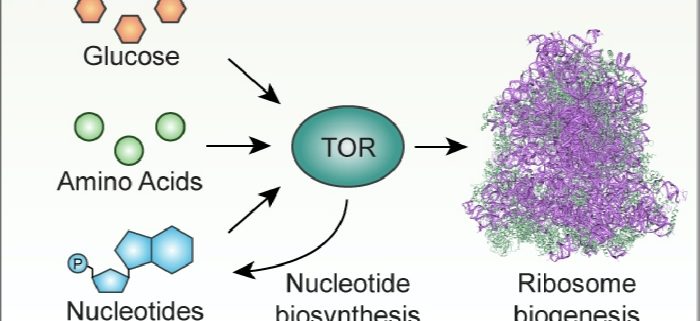
Background: TARGET OF RAPAMYCIN (TOR) is a highly conserved protein kinase that acts as a master regulator of cellular metabolism. In humans and yeast, TOR senses nutrients to promote growth only when the necessary life “building blocks”, especially carbohydrates and amino acids, are available. Recently, researchers in the biomedical field discovered that human TOR also senses the levels of nucleotides, the building blocks of RNA and DNA. However, they did not check whether TOR might also sense nucleotide levels in other species. In fact, much less is known about how TOR senses nutrients and coordinates metabolism in plants.
Question: What nutrients or signals does TOR sense in plants? Are these mechanisms shared across plants, yeast, and animals?
Findings: We conducted a genetic screen to identify pathways that activate TOR. This screen led to the discovery that disrupting nucleotide biosynthesis causes TOR to become less active in two model plant species, Arabidopsis thaliana and Nicotiana benthamiana. Most of the cellular pool of nucleotides is used to synthesize ribosomes, the machines that translate messenger RNAs into functional proteins, so we hypothesized that TOR might sense nucleotide availability to coordinate ribosome biogenesis. Indeed, we showed that when nucleotide biosynthesis and TOR activity are disrupted, ribosome synthesis was repressed. Therefore, we propose that a major function for TOR in plants is to sense when nutrients are available to support ribosome biosynthesis, and thus to sustain mRNA translation.
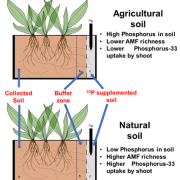
Next steps: Phosphorus is an essential macronutrient for plants, but is not generally abundant in agricultural soils, which limits how well crops grow in more than 30% of global farmland. Most of the phosphorus in plants is used to make nucleotides that are then incorporated into ribosomal RNAs. It therefore comes to reason that plants must somehow sense and incorporate the amount of phosphorus available in their environment when making new ribosomes. By understanding precisely how TOR monitors nucleotide levels, we may discover new genetic targets for breeding plants that use phosphorus more efficiently.
Michael Busche, M. Regina Scarpin, Robert Hnasko, Jacob O. Brunkard (2021). TOR coordinates nucleotide availability with ribosome biogenesis in plants. Plant Cell, https://doi.org/10.1093/plcell/koab043