Plant-herbivore chemical communication decoded (Science)
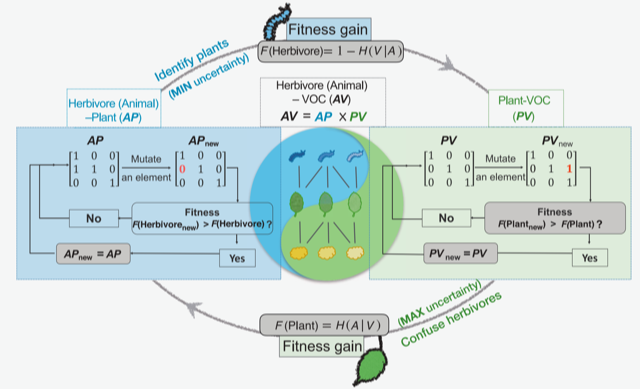 Chemical communications between species are prevalent in nature. For instance, herbivorous insects can spot their host plants by sensing volatile organic compounds (VOCs) emitted by plants. Despite our knowledge about interactions between individual plant and herbivore species, little is known about how chemical communications are structured and maintained in the ecological context. Zu et al. hypothesized that chemical communications between plants and herbivores are shaped by conflicting optimization processes; plants try to decrease the ability of herbivores to detect the right host (in other words, the herbivores’ decoding efficiency) by changing the suite of VOCs they produce. At the same time, herbivores try to increase their decoding efficiency to correctly identify edible plants. In this model, herbivore fitness is proportional to the decoding efficiency, and plant fitness is negatively proportional to it. To validate the model, the authors conducted fieldwork in a tropical dry forest, in which they collected larvae found on plant leaves and sampled VOCs from individual plant species. They then created matrices of plant-herbivore association and plant-VOC association; these two can be combined to generate herbivore-VOC association matrices, which define the decoding efficiency of herbivores. The authors showed that the simulation based on their model could explain key parameters, including the fitness of plants and herbivores observed in the field study. The model suggested that the conflicting optimization process (“information arms race”) drives plants to produce VOCs shared among other plant species to confuse herbivores at the community level, and this potentially pressures herbivores to specialize on a small number of plant species. This study proposes a theoretical framework explaining critical drivers of the formation and maintenance of multi-trophic interactions. (Summary by Tatsuya Nobori @nobolly) Science 10.1126/science.
Chemical communications between species are prevalent in nature. For instance, herbivorous insects can spot their host plants by sensing volatile organic compounds (VOCs) emitted by plants. Despite our knowledge about interactions between individual plant and herbivore species, little is known about how chemical communications are structured and maintained in the ecological context. Zu et al. hypothesized that chemical communications between plants and herbivores are shaped by conflicting optimization processes; plants try to decrease the ability of herbivores to detect the right host (in other words, the herbivores’ decoding efficiency) by changing the suite of VOCs they produce. At the same time, herbivores try to increase their decoding efficiency to correctly identify edible plants. In this model, herbivore fitness is proportional to the decoding efficiency, and plant fitness is negatively proportional to it. To validate the model, the authors conducted fieldwork in a tropical dry forest, in which they collected larvae found on plant leaves and sampled VOCs from individual plant species. They then created matrices of plant-herbivore association and plant-VOC association; these two can be combined to generate herbivore-VOC association matrices, which define the decoding efficiency of herbivores. The authors showed that the simulation based on their model could explain key parameters, including the fitness of plants and herbivores observed in the field study. The model suggested that the conflicting optimization process (“information arms race”) drives plants to produce VOCs shared among other plant species to confuse herbivores at the community level, and this potentially pressures herbivores to specialize on a small number of plant species. This study proposes a theoretical framework explaining critical drivers of the formation and maintenance of multi-trophic interactions. (Summary by Tatsuya Nobori @nobolly) Science 10.1126/science.