NIT proteins regulate rice root plasticity in response to nitrate and ammonium
Lena Maria Müller
Boyce Thompson Institute, Ithaca, NY, USA
Nitrogen (N) is a critical element in plant nutrition and an important building block of amino acids, nucleic acids, and secondary metabolites. Roots take up N mainly in the form of nitrate (NO3–) and ammonium (NH4+). To feed a growing human population and maintain plant growth and crop productivity, most agricultural fields are fertilized with high amounts of N fertilizer. However, excess N fertilizer can leach into the groundwater and pose serious threats to the environment, such as eutrophication of aquatic ecosystems. To achieve more sustainable agriculture, we must reduce N fertilizer usage while maintaining high crop yields. A pivotal prerequisite for achieving these goals is to better understand plant physiological responses to this essential nutrient (Vidal et al., 2020).
NO3– and NH4+ are not only essential nutrients, but are also signaling molecules that influence large-scale transcriptomic changes and developmental adaptations in plants (Liu and Wirén, 2017; Sun et al., 2017). As NO3– and NH4+ are not homogenously distributed in most soils, root systems must dynamically respond to local nutrient availabilities. The root adaptive response to N deprivation or supply is characterized by (i) transcriptional regulation of NO3– or NH4+ uptake systems and (ii) extensive root system remodeling to improve N foraging in the soil (Jia and Wirén, 2020). N-mediated regulation of root system plasticity is complex: high overall N supply as well as severe N starvation inhibit primary root elongation and lateral root development, while low to moderate N supply or heterogeneously distributed N promote primary root growth as well as lateral root branching and elongation, increasing root system N foraging capacities.
Roots acquire NO3– via various low- and high-affinity transporters that belong to different protein families; one such family is the nitrate transporter NRT2. In Oryza sativa, several members of the OsNRT2 protein family were shown to interact with a protein of the NITRATE ASSIMILATION RELATED family (OsNAR2.1) (Yan et al., 2011). OsNAR2.1 itself has no transport capacity, but was reported to be critical for OsNRT2-mediated NO3– uptake. In addition, OsNAR2.1 influences the distribution of auxin in the roots (Huang et al., 2015). Auxin is a known regulator of root system plasticity, and a tight interplay of this phytohormone and NO3–-signaling is well documented (Zhang et al., 1999). However, the mechanistic connection of OsNAR2.1 function, NO3– -uptake, and auxin signaling were unclear.
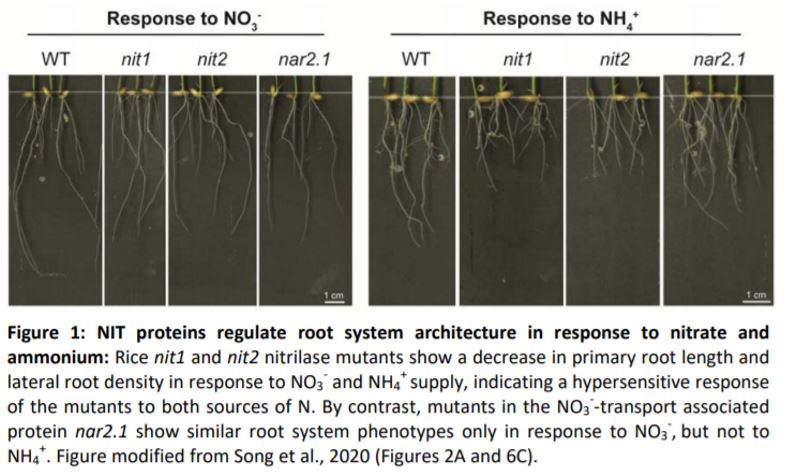 In this issue of Plant Physiology, Song et al. report the identification of two nitrilases, OsNIT1 and OsNIT2, as binding partners of OsNAR2.1. OsNIT1 and OsNIT2, as well as OsNAR2.1, are mainly expressed in the tips of lateral roots but also in mature roots, and expression of all three genes is induced after root treatment with NO3–. Using single and double mutant analyses, the authors established that OsNIT1, OsNIT2, and OsNAR2.1 confer partially overlapping but also separate functions: all three genes are involved in shaping root architecture in response to NO3– by promoting primary root growth and increasing lateral root density (Figure 1). However, in contrast to nar2.1 (Huang et al., 2015), nit1 or nit2 knockout mutants were not affected in uptake of radiolabelled 15NO3–. This indicates that OsNIT1 and OsNIT2 have a function in signaling during root system adaptation to fluctuating NO3– supply rather than directly contributing to NO3– uptake, whereas OsNAR2.1 contributes to both processes.
In this issue of Plant Physiology, Song et al. report the identification of two nitrilases, OsNIT1 and OsNIT2, as binding partners of OsNAR2.1. OsNIT1 and OsNIT2, as well as OsNAR2.1, are mainly expressed in the tips of lateral roots but also in mature roots, and expression of all three genes is induced after root treatment with NO3–. Using single and double mutant analyses, the authors established that OsNIT1, OsNIT2, and OsNAR2.1 confer partially overlapping but also separate functions: all three genes are involved in shaping root architecture in response to NO3– by promoting primary root growth and increasing lateral root density (Figure 1). However, in contrast to nar2.1 (Huang et al., 2015), nit1 or nit2 knockout mutants were not affected in uptake of radiolabelled 15NO3–. This indicates that OsNIT1 and OsNIT2 have a function in signaling during root system adaptation to fluctuating NO3– supply rather than directly contributing to NO3– uptake, whereas OsNAR2.1 contributes to both processes.
In other plant species, some nitrilases have been implicated in the conversion of indole-3-acetonitrile (IAN) to indole-3-acetic acid (IAA), the most abundant form of auxin (Enders and Strader, 2015). Therefore, Song et al. tested if the OsNIT1 and OsNIT2 signaling functions are related to auxin biosynthesis or transport. While the authors did not detect changes in free IAA content in nit1 or nit2 mutant roots relative to the wild type—which could possibly be explained by the existence of NIT-independent auxin biosynthetic pathways (Enders and Strader, 2015)—in vitro studies suggested that the nitrilase OsNIT1 is indeed capable of converting IAN to IAA. This enzymatic activity was even stronger when OsNIT2 and OsNAR2.1 were present in the same reaction, indicating that the protein complex might be involved in auxin biosynthesis in response to NO3– in roots. In addition, OsNIT1 and OsNIT2 affected the distribution of auxin in roots: by locally applying radiolabelled [3H+]-IAA to the root–shoot junction and subsequently measuring [3H+]-IAA in the root tips, Song et al. demonstrated that nit1 and nit2 mutants are defective in root acropetal (toward the root tip) auxin movement. Thus, a concerted action of OsNAR2.1, OsNIT1, and OsNIT2 possibly couples NO3– uptake with auxin biosynthesis and/or distribution, which in turn could explain the effect of these genes on primary and lateral root growth in response to NO3–.
Although NH4+ is taken up by distinct OsNAR2.1-independent uptake systems that involve permeases of the AMT protein family (Gazzarrini et al., 1999), Song et al. found that, in addition to NO3–, root expression of OsNIT1 and OsNIT2 was also induced by NH4+. Similar to their NO3– experiments, Song et al. report that OsNIT1 and OsNIT2 influence root system architecture and auxin distribution also in the response to NH4+ (Figure 1), indicating that the two nitrilases could be members of both the NO3– and NH4+ signaling pathways. By contrast, OsNAR2.1 functions specifically in NO3– signaling and uptake, but is not regulated by NH4+ (Huang et al., 2015). The authors hypothesize that OsNAR2.1 likely increases OsNIT function specifically in response to NO3–. It remains to be investigated if similar reinforcing factors exist that increase nitrilase function in response to NH4+. Thus, the NIT proteins could be a critical intersection of both NO3– and NH4+ signaling pathways, ensuring adaptive root responses to optimize N acquisition regardless of the uptake form. These findings bring us one step closer to understanding rice N nutrition, which could potentially allow us to generate crops with increased N uptake capacities for a more sustainable agriculture with reduced N fertilizer usage.
References
Enders TA, Strader LC (2015) Auxin activity: Past, present, and future. American Journal of Botany 102: 180–196
Gazzarrini S, Lejay L, Gojon A, Ninnemann O, Frommer WB, Wirén von N (1999) Three functional transporters for constitutive, diurnally regulated, and starvation-induced uptake of ammonium into Arabidopsis roots. The Plant Cell 11: 937–948
Huang S, Chen S, Liang Z, Zhang C, Yan M, Chen J, Xu G, Fan X, Zhang Y (2015) Knockdown of the partner protein OsNAR2.1 for high-affinity nitrate transport represses lateral root formation in a nitrate-dependent manner. Sci Rep 5: 18192–12
Jia Z, Wirén NV (2020) Signaling pathways underlying nitrogen-dependent changes in root system architecture: from model to crop species. Journal of experimental botany 80: 1
Liu Y, Wirén von N (2017) Ammonium as a signal for physiological and morphological responses in plants. Journal of experimental botany 68: 2581–2592
Sun C-H, Yu J-Q, Hu D-G (2017) Nitrate: A Crucial Signal during Lateral Roots Development. Front Plant Sci 8: 485
Vidal EA, Alvarez JM, Araus V, Riveras E, Brooks M, Krouk G, Ruffel S, Lejay L, Crawford N, Coruzzi GM, et al (2020) Nitrate 2020: Thirty years from transport to signaling networks. The Plant cell tpc.00748.2019
Yan M, Fan X, Feng H, Miller AJ, Shen Q, Xu G (2011) Rice OsNAR2.1 interacts with OsNRT2.1, OsNRT2.2 and OsNRT2.3a nitrate transporters to provide uptake over high and low concentration ranges. Plant Cell Environ 34: 1360–1372
Zhang H, Jennings A, Barlow PW, Forde BG (1999) Dual pathways for regulation of root branching by nitrate. Proceedings of the National Academy of Sciences 96: 6529–6534



