It’s All in the Neighborhood: SHORTROOT-mediated Intercellular Signals Coordinate Phloem Development in the Root
In all multicellular organisms, tissue patterning and organogenesis are governed by asymmetric cell divisions (ACDs) and the cellular exchange of positional cues. Plants, unlike animals, undergo post-embryonic organogenesis due to the maintenance of stem cells in specialized niches. One such niche, the root apical meristem, is found near the tip of the root. Many studies over the last three decades have used the root apical meristem as a model to study the molecular regulation of post-embryonic tissue patterning. Kim and Zhou et al. (2020) uncover a new role for SHORTROOT (SHR), a mobile transcription factor, in coordinating phloem tissue development in the root.
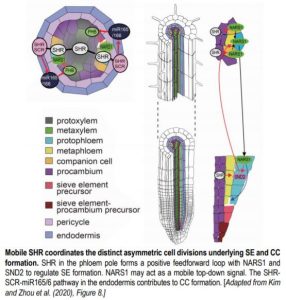
Phloem is important for the long-distance transport of nutrients from source to sink. There are two major phloem cell types: companion cells (CCs; orange in Figure) and proto- and meta-phloem sieve elements (SEs; yellow and light blue in Figure). Although the development of CCs and SEs is coordinated, each cell type is patterned via distinct ACDs of different precursor cells. Molecular regulators of phloem development, including both local and long-distance signals, have recently been reported (Blob et al., 2018). However, the mechanism that coordinates these two distinct ACDs in space and time is unknown.
 Given its functions in xylem and endodermis development (Carlsbecker et al., 2010; Drapek et al., 2017), Kim and Zhou et al. (2020) hypothesized that SHR may also be involved in phloem development. Genetic data indeed suggest that a loss of SHR function compromises cell division activities required for CC and SE formation. Previous work revealed that SHR in the endodermis and quiescent center activates another transcription factor, SCARECROW (SCR), and together the two initiate the expression of mobile microRNA165/166 (miR165/6) to pattern xylem vessels (Carlsbecker et al., 2010). Here, genetic experiments suggest that the SHR-SCR-miR165/6 pathway contributes to the control of procambial cell proliferation, including ACD for CCs, but does not contribute to ACD for SE formation.
Given its functions in xylem and endodermis development (Carlsbecker et al., 2010; Drapek et al., 2017), Kim and Zhou et al. (2020) hypothesized that SHR may also be involved in phloem development. Genetic data indeed suggest that a loss of SHR function compromises cell division activities required for CC and SE formation. Previous work revealed that SHR in the endodermis and quiescent center activates another transcription factor, SCARECROW (SCR), and together the two initiate the expression of mobile microRNA165/166 (miR165/6) to pattern xylem vessels (Carlsbecker et al., 2010). Here, genetic experiments suggest that the SHR-SCR-miR165/6 pathway contributes to the control of procambial cell proliferation, including ACD for CCs, but does not contribute to ACD for SE formation.
To determine what controls SE formation, the authors hypothesized the involvement of SHR in the stele itself. Phenotypes resulting from the expression of GFP-tagged SHR either in the stele or specifically in the phloem of shr mutants, together with a complementary experiment to block SHR mobility, suggest that SHR moving into the phloem pole promotes ACD for SE formation. Based on this result, the authors hypothesized that SHR regulates the expression of additional transcription factors specifically in the phloem. A combination of phloem-enriched genes identified from existing expression data, a comparative microarray experiment, and a time course induction assay identified five transcription factor candidates with increased expression in response to SHR induction. Chromatin immunoprecipitation experiments indicated that SHR can bind to the promoters of two candidates, NAC-REGULATED SEED MORPHOLOGY 1 (NARS1) and SECONDARY WALL-ASSOCIATED NAC DOMAIN PROTEIN 2 (SND2). Genetic analyses suggest that NARS1 regulates the ACD of the phloem SE precursor as a downstream target of SHR.
NARS1 is upregulated soon after SHR induction but SND2 expression is detected much later, suggesting that SND2 induction requires other SHR-dependent factors. From a comprehensive set of genetic experiments, the authors conclude that SHR, NARS1, and SND2 form a positive feedforward loop to regulate phloem SE development (see Figure). Interestingly, although data suggest that NARS1 acts locally to control the ACD leading to SE development, NARS1 expressed in differentiated phloem CCs was shown to activate SND2 in the young SE cells of the meristem. NARS1 may therefore act as a mobile signal, though additional data is needed to test NARS1 protein mobility.
Taken together, this work produces a model for the coordination of phloem development based on SHR movement. SHR moving into the endodermis regulates the ACD that produces the CC while SHR that moves into the phloem pole is primarily responsible for the ACD that forms the SE. Future work will unravel precisely how NARS1 regulates ACDs of the SE precursor.
Rachel Shahan
Department of Biology, Duke University
Durham, North Carolina
ORCID: 0000‐0001‐7702‐9432
REFERENCES
Blob B, Heo JO, and Helariutta Y. (2018). Phloem differentiation: an integrative model for cell specification. J. Plant Res. 131, 31-36.
Carlsbecker A, Lee J-Y, Roberts CJ, Dettmer J, Lehesranta S, Zhou J, Lindgren O, Moreno-Risueno MA, Vatén A, Thitamadee S, Campilho A, Sebastian J, Bowman JL, Helariutta Y, and Benfey PN. (2010). Cell signalling by microRNA165/6 directs gene dose-dependent root cell fate. Nature. 465, 316-321.
Drapek C, Sparks EE, and Benfey PN. (2017). Uncovering Gene Regulatory Networks Controlling Plant Cell Differentiation. Trends Genet. 33, 529–539.
Kim H, Zhou J, Kumar D, Jang G, Ryu KH, Sebastian J, Miyashima S, Helariutta Y, and Lee J-Y (2020). SHORTROOT-mediated intercellular signals that coordinate the phloem development process in Arabidopsis roots. DOI: https://doi.org/10.1105/tpc.19.00455



