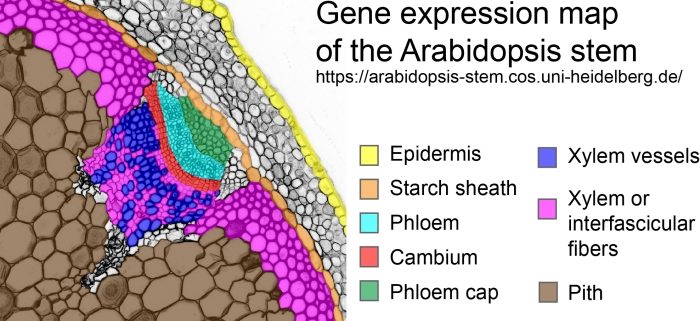
Gene Expression Map of the Arabidopsis Stem
Shi et al. provide a high-resolution gene expression map of the Arabidopsis stem. The Plant Cell (2021) http://bit.ly/3pgWm4G
By Dongbo Shi and Thomas Greb
Bckground: Multicellular organisms are usually built of numerous different tissues with very special functions. These functions depend on unique sets of gene activities. Therefore, obtaining information about which gene is active in a certain tissue is vital for understanding the complexity of body organization. Plant stems transport water, sugars, and many signaling molecules and play fundamental roles in determining shoot architecture and plant growth overall. These functions are fulfilled by very specialized cells, such as water-transporting vessel elements and nutrient-transporting phloem. Importantly, since plant cells are surrounded by connecting cell walls, tissue isolation is often a bottleneck for obtaining information on gene activity; mature organs such as the stem are especially difficult to analyze.
Question: In this work, we explored whether we could provide high-resolution gene expression maps of mature plant organs such as the stem by analyzing mRNA populations found in nuclei, which are much easier to harvest than whole cells.
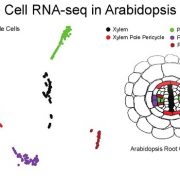
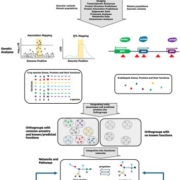
Findings: We labeled nuclei in individual tissues of Arabidopsis thaliana stems using fluorescent proteins by expressing these proteins under the control of gene promoters specifically active in cells such as vessel elements or phloem cells. After fluorescence-based nucleus sorting and collecting mRNA from fluorescent nuclei of each tissue, we were able to characterize mRNA profiles indicating the activity of each gene in the genome. By validating the discovered patterns of a large number of gene activities in stems, we indeed showed that gene activity in stems could be estimated by investigating nuclear mRNA. Using this technique combined with laser-capture microdissection, another method for isolating cellular material from distinct tissues, we generated a comprehensive gene expression map of the Arabidopsis stem. Based on sequence analysis of the promoter regions of the identified genes, we also identified candidate tissue-specific gene regulators.
Next steps: Our datasets will be beneficial for researchers generating hypotheses towards the spatial organization of physiological processes in plant stems. We plan to apply our methods to other plant organs, use multi-color labeling of nuclei to further increase spatial resolution, and perform profiling of single nuclei.
Dongbo Shi, Virginie Jouannet, Javier Agustí, Verena Kaul, Victor Levitsky, Pablo Sanchez, Victoria V Mironova, Thomas Greb (2021). Tissue-specific transcriptome profiling of the Arabidopsis inflorescence stem reveals local cellular signatures. Plant Cell http://bit.ly/3pgWm4G