Evolution of C4 photosynthesis in the Cleomaceae
Background: The Cleomaceae is the sister family to the Brassicaceae (including the model species Arabidopsis and Brassica crops). The Cleomaceae contains species with different types of photosynthesis, including C3, C4 and C3-C4 intermediate plants. As the Brassicaceae family does not have a true C4 species, the Cleomaceae serves as a valuable model system for photosynthesis research that aims to improve crops. The Cleomaceae also includes several economically important leafy, medicinal and ornamental plants. Despite its scientific and economical importance, few genetic and genomic resources exist for the Cleomaceae.
Question: How did the Cleomaceae family evolve since its divergence from the Brassicaceae? What factors contributed to the evolution of C4 photosynthesis in Cleomaceae?
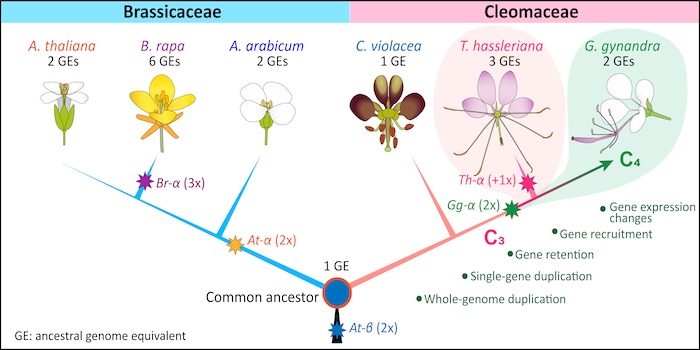
Findings: We generated a reference genome for the C4 species G. gynandra that facilitates comparative genomics with its C3 relative, Tarenaya hassleriana, to elucidate the family polyploidy history and evolution of C4 photosynthesis in the Cleomaceae. These species evolved through step-wise ancient polyploidy events, in which a whole-genome duplication event (Gg-α) occurred first, followed by an addition of a third genome (Th-α, +1x) to T. hassleriana but not to G. gynandra. The evolution of C4 photosynthesis in the Cleomaceae resulted from a series of processes, including differential duplication, retention, recruitment and expression modification of C4-related genes. This led to the preferential expression of these genes in leaf mesophyll or bundle-sheath cells depending on their functions.
Next steps: Future efforts will focus on developing genomic resources for species of different photosynthesis types in the Cleomaceae. This will allow a more systematic analysis of the family history and trait evolution. It will also facilitate the study of important gene families related to plant physiological and anatomical changes involved in the transition from C3 to C4 photosynthesis. This can help to engineer C4 photosynthesis into non-C4 crops.
Reference:
Nam V. Hoang, E. O. Deedi Sogbohossou, Wei Xiong, Conor J. C. Simpson, Pallavi Singh, Nora Walden, Erik van den Bergh, Frank F. M. Becker, Zheng Li, Xin-Guang Zhu, Andrea Brautigam, Andreas P. M. Weber, Jan C. van Haarst, Elio G. W. M. Schijlen, Prasad S. Hendre, Allen Van Deynze, Enoch G. Achigan-Dako, Julian M. Hibberd, M. Eric Schranz (2023). The Gynandropsis gynandra genome provides insights into whole-genome duplications and the evolution of C4 photosynthesis in Cleomaceae. https://doi.org/10.1093/plcell/koad018



