Collinear Chromosomes and Shifting Centromeres in the Arabideae
For such an important structure, the centromere certainly is hard to pin down, particularly in regard to the relationship between its sequence composition and epigenetic state and its functions in chromosome pairing and kinetochore formation. In addition to undergoing rapid sequence evolution, centromeres can undergo changes in location over evolutionary time. In many cases, this change involves large-scale chromosome rearrangements; in other cases, the genes on the chromosomes remain collinear, but the centromere position shifts, forming an Evolutionary New Centromere (ENC, reviewed in Schubert, 2018).
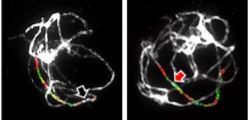
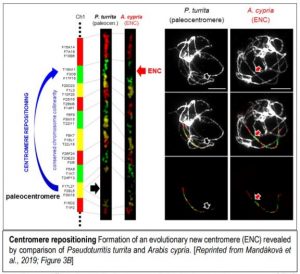 In a recent paper in The Plant Cell, Mandáková et al. (2020) examine whether centromere position has changed between homeologous chromosomes of different species and genera within the Arabideae, a large and diverse clade within the Brassicaceae. Their interest was piqued by observations of changes in centromere position in the sequenced genome of Alpine rockcress (Arabis alpina) and they then extended this analysis to multiple species in the Arabideae, starting with Pseudoturritis turrita, which has the most-ancestral position in the Arabideae––that is, it’s the earliest branch off of the phylogenetic tree. To examine the genome structure of these species, they first used chromosome painting––in situ hybridization with chromosome-specific bacterial artificial chromosomes (BACs) from Arabidopsis thaliana––to identify 22 conserved genomic blocks of the eight chromosomes. Comparison of P. turrita with other members of the Arabideae showed that chromosomes had undergone translocations in different lineages but, for six homoeologs, the gene order remained collinear––and for five of these six, the centromere position had changed, indicating the formation of an ENC (see Figure). The authors additionally identified tandem repeats that localized to the centromere in various Arabideae species. Collinearity comparison between the sequenced genomes of A. alpina and Arabidopsis lyrata confirmed the results from chromosome painting. Moreover, this sequence comparison showed that the pericentromeric regions of ENCs and centromeres that have remained in the same place did not differ substantially in repeat content, DNA methylation, or size.
In a recent paper in The Plant Cell, Mandáková et al. (2020) examine whether centromere position has changed between homeologous chromosomes of different species and genera within the Arabideae, a large and diverse clade within the Brassicaceae. Their interest was piqued by observations of changes in centromere position in the sequenced genome of Alpine rockcress (Arabis alpina) and they then extended this analysis to multiple species in the Arabideae, starting with Pseudoturritis turrita, which has the most-ancestral position in the Arabideae––that is, it’s the earliest branch off of the phylogenetic tree. To examine the genome structure of these species, they first used chromosome painting––in situ hybridization with chromosome-specific bacterial artificial chromosomes (BACs) from Arabidopsis thaliana––to identify 22 conserved genomic blocks of the eight chromosomes. Comparison of P. turrita with other members of the Arabideae showed that chromosomes had undergone translocations in different lineages but, for six homoeologs, the gene order remained collinear––and for five of these six, the centromere position had changed, indicating the formation of an ENC (see Figure). The authors additionally identified tandem repeats that localized to the centromere in various Arabideae species. Collinearity comparison between the sequenced genomes of A. alpina and Arabidopsis lyrata confirmed the results from chromosome painting. Moreover, this sequence comparison showed that the pericentromeric regions of ENCs and centromeres that have remained in the same place did not differ substantially in repeat content, DNA methylation, or size.
Therefore, the authors have shown that members of the Arabideae tend to have similar numbers of chromosomes and the genes on these chromosomes tend to remain in the same order, but the centromeres on many of these chromosomes have changed in position. Changes in centromere position have major implications for speciation, chromosome pairing in polyploid plants, and for plant evolution in general, but the mechanisms remain unclear with many intriguing questions for future research. For example, does the re-positioning occur by gradual centromere movement (sliding), or does the centromere fade away in one spot and form in a new position (seeding)? If a new centromere is seeded at a different location, when (and how!) does the old centromere hand over control to the new centromere, thereby avoiding acentric or dicentric chromosomes? In their comprehensive Discussion section, Mandáková et al. (2020) examine the implications of their data for many of these questions, such as whether ENCs tend to form in regions with specific characteristics and the relationship between the emergence of ENCs and speciation in the Arabideae. Additional, in-depth studies (particularly as more genomes are sequenced in this tribe) will surely shed more light on the outstanding, exciting questions raised by this study.
Science Editor, Jennifer Mach
http://orcid.org/0000-0002-1141-6306
REFERENCES
Mandáková, T., Hloušková, P., Koch, M.A., and Lysak, M.A. (2020). Genome Evolution in Arabideae was Marked by Frequent Centromere Repositioning. Plant Cell DOI: https://doi.org/10.1105/tpc.19.00557.
Schubert, I. (2018) What is behind “centromere repositioning”?. Chromosoma 127: 229-234