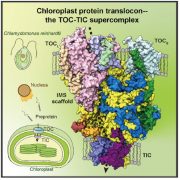
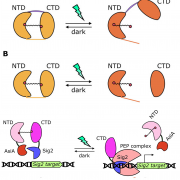
Chloroplast protein import determines plant proteostasis and retrograde signaling (bioRxiv)
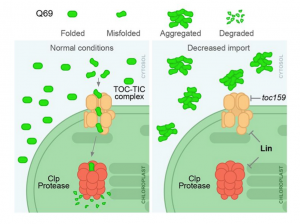 The neurological condition Huntington’s disease is caused by mutated versions of the Huntingtin gene that encode expanded stretches of polyglutamine repeats, resulting in protein aggregation. However, despite the widespread presence of polyglutamine repeat regions in plant proteomes, similar pathologies have yet to be observed. To investigate this paradox, Llamas and colleagues expressed aggregation-prone fragments of the human Huntingtin protein, Q28 and Q69, in Arabidopsis thaliana plants. Amazingly the protein did not aggregate and as a result the plants remained healthy. Further protein interaction studies revealed that Q28 and Q69 are imported into the chloroplast and degraded by the stromal processing peptidase (SPP) protein. The researchers then co-expressed a version of the Arabidopsis SPP protein in a human cell line also expressing a polyglutamine protein. Remarkably these co-expression cells showed reduced aggregation of the protein, suggesting a potential therapeutic avenue for the treatment of these diseases in humans. The group also showed that inhibition of chloroplast protein import not only led to the aggregation of Q69, but also of an endogenous plant polyglutamine containing protein, plastid casein kinase 2. Under these conditions, the protein forms cytosolic aggregates and accumulates in the nucleus, suggesting a potential role for polyglutamate-containing proteins in plant organellar retrograde signaling. (Summary by Rory Burke @rorby95) bioRxiv 10.1101/2022.03.19.484971 (And don’t miss the animated Tweetorial created by the first author).
The neurological condition Huntington’s disease is caused by mutated versions of the Huntingtin gene that encode expanded stretches of polyglutamine repeats, resulting in protein aggregation. However, despite the widespread presence of polyglutamine repeat regions in plant proteomes, similar pathologies have yet to be observed. To investigate this paradox, Llamas and colleagues expressed aggregation-prone fragments of the human Huntingtin protein, Q28 and Q69, in Arabidopsis thaliana plants. Amazingly the protein did not aggregate and as a result the plants remained healthy. Further protein interaction studies revealed that Q28 and Q69 are imported into the chloroplast and degraded by the stromal processing peptidase (SPP) protein. The researchers then co-expressed a version of the Arabidopsis SPP protein in a human cell line also expressing a polyglutamine protein. Remarkably these co-expression cells showed reduced aggregation of the protein, suggesting a potential therapeutic avenue for the treatment of these diseases in humans. The group also showed that inhibition of chloroplast protein import not only led to the aggregation of Q69, but also of an endogenous plant polyglutamine containing protein, plastid casein kinase 2. Under these conditions, the protein forms cytosolic aggregates and accumulates in the nucleus, suggesting a potential role for polyglutamate-containing proteins in plant organellar retrograde signaling. (Summary by Rory Burke @rorby95) bioRxiv 10.1101/2022.03.19.484971 (And don’t miss the animated Tweetorial created by the first author).