Changing gears in salt stress tolerance: AtCAMTA6’s role in transcriptional regulation of ion transport
Salt stress affects plant growth and development at all stages, reducing germination, and compromising seedling establishment, transpiration, and root growth (Munns and Tester, 2008). At later developmental stages, the overaccumulation of salt ions in the shoot disrupts photosynthesis and other vital processes, oftentimes resulting in premature leaf senescence. The release of vacuolar Ca2+ into the cytosol occurs early after salt stress exposure and is essential for propagation of stress signals throughout the plant (Choi et al., 2014). One of the best characterized Ca2+ sensors within the plant cell is calmodulin (CaM) (Virdi et al., 2015), which undergoes a conformational change upon Ca2+ binding, thereby changing its affinity for interacting proteins. CaM can affect general transcription levels either indirectly, through interactions with protein kinases, or directly, by interacting with transcription factors. Six CaM-binding transcriptional activators (CAMTAs) identified in Arabidopsis thaliana (Bouché et al., 2002) show induced transcription levels in response to biotic and abiotic stress (Yang and Poovaiah, 2002; Doherty et al., 2009; Pandey et al., 2013). CAMTAs have conserved protein structures, containing the CaM-binding IQ motifs towards the C-terminus and DNA-binding domains at the N-terminal end (Finkler et al., 2007). While the majority of the CAMTAs are well studied, the function of CAMTA6 remained elusive until recently.
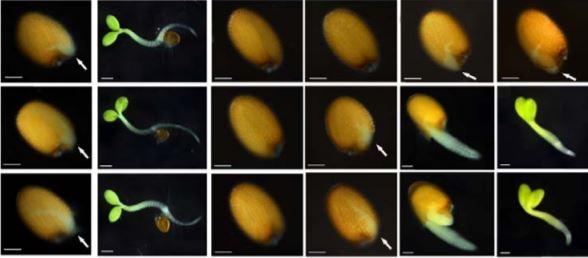
In this issue of Plant Physiology, Shkolnik (et al., 2019) describe the role of AtCAMTA6 in salt tolerance during seedling germination. The group observed that camta6 mutants showed an increased germination rate under salt stress conditions as severe as 200 mM NaCl. This increase in germination could be explained by reduced sodium uptake by the germinating seed, observed for two independent camta6 alleles. Interestingly, Shkolnik (et al., 2019) observed that decreased sodium accumulation corresponded to the repressed expression of High-affinity K+ Transporter (AtHKT1) known to import sodium ions into the root pericycle from the vasculature, thereby limiting sodium accumulation in the shoot (Mäser et al., 2002; Møller et al., 2009). The levels of AtHKT1 were not induced in response to salt stress in camta6 lines, and AtHKT1 expression was also limited to the radicle and excluded from developing cotyledons.
AtHKT1 is not the only gene affected by AtCAMTA6, as the transcriptomic study of camta6 lines revealed that CAMTA is responsible for more than half of the salt-stress induced transcriptomic changes at the germination stage. Future developments on how CAMTA6 regulates the transcriptome will shed more light on its direct and indirect targets as well as the integration of calcium and ABA signaling during salt stress.
Another exciting aspect of the work described by Shkolnik (et al., 2019) is that while camta6 seedlings outperformed wild-type lines during germination under salt stress, reduced salt stress tolerance was observed for the mutants at later developmental stages. This phenomenon could be explained by camta6 lines’ inability to induce AtHKT1 expression. Enhanced HKT1 expression improves salinity tolerance in older plants (Møller et al., 2009). In younger seedlings, by contrast, high HKT1 expression results in reduced lateral root development and increased salt stress sensitivity (Julkowska et al., 2017). While HKT1 is one of many genes affected by CAMTA6, this study suggests that ion sequestration mechanism, involving AtHKT1 and AtCAMTA6, reduces salinity tolerance at the early developmental stage but is necessary later in plant development. In the future, increasing our understanding of the upstream regulation of CAMTA6 will provide a better understanding of this developmental switch in preference for salt tolerance strategy, and could be used in creating more dynamic salt tolerance mechanisms in crop species.
References:
Bouché N, Scharlat A, Snedden W, Bouchez D, Fromm H (2002) A novel family of calmodulin-binding transcription activators in multicellular organisms. J Biol Chem 277: 21851–21861
Choi W-G, Toyota M, Kim S-H, Hilleary R, Gilroy S (2014) Salt stress-induced Ca2+ waves are associated with rapid, long-distance root-to-shoot signaling in plants. Proc Natl Acad Sci U S A 111: 6497–6502
Doherty CJ, Van Buskirk HA, Myers SJ, Thomashow MF (2009) Roles for Arabidopsis CAMTA transcription factors in cold-regulated gene expression and freezing tolerance. Plant Cell 21: 972–984
Finkler A, Ashery-Padan R, Fromm H (2007) CAMTAs: calmodulin-binding transcription activators from plants to human. FEBS Lett 581: 3893–3898
Julkowska MM, Koevoets IT, Mol S, Hoefsloot H, Feron R, Tester MA, Keurentjes JJB, Korte A, Haring MA, de Boer G-J, et al (2017) Genetic Components of Root Architecture Remodeling in Response to Salt Stress. Plant Cell 29: 3198–3213
Mäser P, Eckelman B, Vaidyanathan R, Horie T, Fairbairn DJ, Kubo M, Yamagami M, Yamaguchi K, Nishimura M, Uozumi N, et al (2002) Altered shoot/root Na+ distribution and bifurcating salt sensitivity in Arabidopsis by genetic disruption of the Na+ transporter AtHKT1. FEBS Lett 531: 157–161
Møller IS, Gilliham M, Jha D, Mayo GM, Roy SJ (2009) Shoot Na+ exclusion and increased salinity tolerance engineered by cell type–specific alteration of Na+ transport in Arabidopsis. The Plant
Munns R, Tester M (2008) Mechanisms of Salinity Tolerance. Annual Review of Plant Biology 59: 651–681
Shkolnik D, Finkler A, Pasmanik-Chor, M, Fromm H (2019) Calmodulin-binding transcritpion activator 6: a key of Na+ homeostasis during germination. Plant Phys. XX: AA-BB
Pandey N, Ranjan A, Pant P, Tripathi RK, Ateek F, Pandey HP, Patre UV, Sawant SV (2013) CAMTA 1 regulates drought responses in Arabidopsis thaliana. BMC Genomics 14: 216
Virdi AS, Singh S, Singh P (2015) Abiotic stress responses in plants: roles of calmodulin-regulated proteins. Front Plant Sci 6: 809
Yang T, Poovaiah BW (2002) A calmodulin-binding/CGCG box DNA-binding protein family involved in multiple signaling pathways in plants. J Biol Chem 277: 45049–45058