A Chloroplast tRNA-Modifying Enzyme Functions in Plant Development
Liu, Ren et al. discover how a natural allele of tRNA-modifying GTPase gene PDD leads to pleiotropic developmental defects in rice. Plant Cell https://doi.org/10.1105/tpc.19.00660.
By Hui Liu and Pingli Lu
Fudan University and Henan University
Background: Transfer RNAs (tRNAs) are components of the translational machinery that deliver amino acids to the protein elongation chain by correctly deciphering the genetic code. tRNA modifications are widespread, affecting both the structure and translation of tRNAs. tRNA modifications are catalyzed by tRNA-modifying enzymes, which usually have conserved functions in eukaryotic and prokaryotic organisms. Mutations in genes encoding tRNA-modifying enzymes that function in the cytoplasm and organelles can greatly affect various developmental processes. However, few functional studies have been performed on genes encoding tRNA-modifying enzymes in the chloroplast, an important semiautonomous organelle in flowering plants.
Question: We were eager to discover whether and how tRNA modifications in the chloroplast affect plant development.
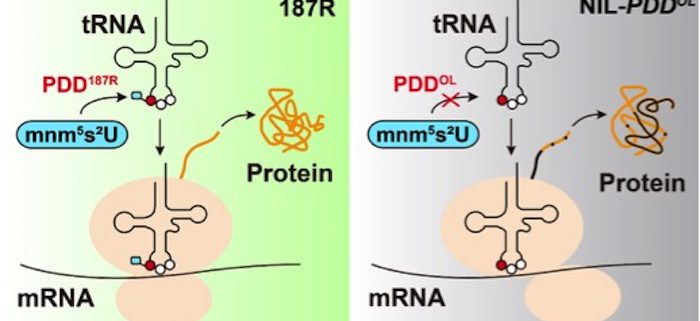
Findings: We identified a natural variation allele of PDD (PLEIOTROPIC DEVELOPMENTAL DEFECTS) from an introgression line (NIL-PDDOL) derived from African wild rice named PDDOL. NIL-PDDOL shows multiple developmental defects. PDD is annotated as a tRNA-modifying GTPase that is predicted to be localized to the chloroplast. Using genetic studies, we revealed that the PDD gene is essential for normal rice development. We obtained several CRISPR-Cas9 edited mutants that survived but died early, indicating that PDDOL is a relatively weak allele, which offered us the opportunity to study its function. PDD mRNA is primarily expressed in green tissues, and PDD is targeted to chloroplasts. PDD can form homodimers and has strong GTPase activity in the wild-type cultivar 187R, whereas PDDOL fails to form homodimers and has weak GTPase activity. We demonstrate that PDD is associated with tRNA modification in chloroplast. The defect of PDD greatly affects the levels of chloroplast proteins involved in both photosynthesis and ribosome biogenesis and alters both chloroplast and nuclear gene expression.
Next steps: The substrate specificity of PDD in chloroplasts is crucial, because the loss of tRNA modification may result in nonfunctional tRNA or tRNA with an aberrant function that consequently produces dysfunctional or deleterious proteins. Identifying the tRNA species that are directly modified by PDD is crucial for further understanding this important biological process.
Hui Liu, Ding Ren, Ling Jiang, Xiaojing Li, Yuan Yao, Limin Mi, Wanli Chen, Aowei Mo, Ning Jiang, Jinshui Yang, Peng Chen, Hong Ma, Xiaojin Luo, and Pingli Lu. (2020). A Natural Variation in PLEIOTROPIC DEVELOPMENTAL DEFECTS Uncovers a Crucial Role for Chloroplast tRNA Modification in Translation and Plant Development. Plant Cell DOI: https://doi.org/10.1105/tpc.19.00660