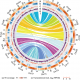
Single-cell RNA sequencing resolves molecular relationships among individual plant cells (Plant Physiol)
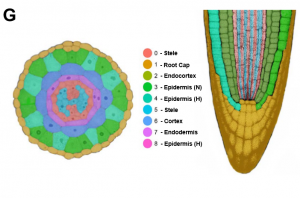 Single cell RNA sequencing (scRNA-seq) has transformed our understanding of gene expression within and amongst individual cells. These techniques have been applied extensively to animal cell populations to discover the development of specific cell lines and identify rare cell types but are not commonly used in plant biology. The study here presents high-throughput scRNA-seq data for Arabidopsis thaliana seedling roots. Analysis of single cell transcriptomic profiles reveals that gene expression from all root tissues are identified improving insights into cell-type differentiation. Further work demonstrates that the approach has the power to recognise rare cell types and differentiate developmental signatures between wild and mutant plants. The paper highlights the untapped potential of scRNA-seq to provide gene expression at single cell resolution which could be applied to other species and plant organs (Summary by Alex Bowles) Plant Physiol. 10.1104/pp.18.01482.
Single cell RNA sequencing (scRNA-seq) has transformed our understanding of gene expression within and amongst individual cells. These techniques have been applied extensively to animal cell populations to discover the development of specific cell lines and identify rare cell types but are not commonly used in plant biology. The study here presents high-throughput scRNA-seq data for Arabidopsis thaliana seedling roots. Analysis of single cell transcriptomic profiles reveals that gene expression from all root tissues are identified improving insights into cell-type differentiation. Further work demonstrates that the approach has the power to recognise rare cell types and differentiate developmental signatures between wild and mutant plants. The paper highlights the untapped potential of scRNA-seq to provide gene expression at single cell resolution which could be applied to other species and plant organs (Summary by Alex Bowles) Plant Physiol. 10.1104/pp.18.01482.