The dark side of the plant: lipid metabolism regulation under starvation
By Corina Mariana Fusari1 & Yariv Brotman2
1Centro de Estudios Fotosintéticos y Bioquímicos (CEFOBI-CONICET-UNR), Suipacha 570, S2000LRJ Rosario, Argentina
2Department of Life Sciences, Ben Gurion University of the Negev, Beersheva, Israel
Background: In the light, plants use photosynthesis to produce sugars and grow, while in the dark, they mainly use starch as the source of energy for maintenance throughout the night. When dark conditions last for a long time, plants consume all the available starch and enter a starvation mode, where they start feeding on lipids to support their metabolism. Although metabolism affects many responses, its genetic regulation is still underexplored. The study of complex traits such as metabolism can be facilitated by combining the use of natural populations and metabolomics. This strategy, called metabolomic-GWAS (Genome Wide Association Studies) enables the identification of genes involved in variation of multiple metabolic features under diverse environmental set-ups.
Question: How is metabolism controlled upon stress? What are the genes acting behind a metabolic response to environmental fluctuations? How did plants evolve to cope with adverse environmental conditions?
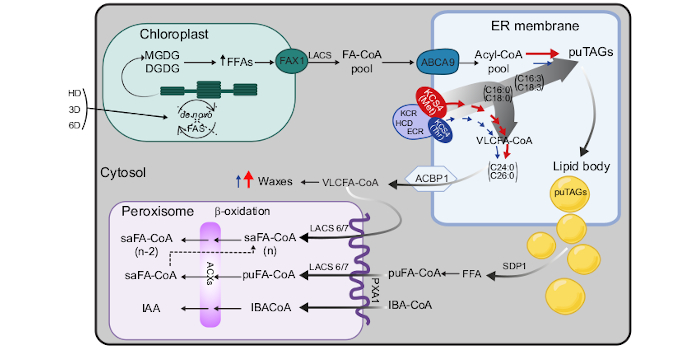
Findings: We were interested in determining the genetic regulation behind the adjustment in lipid metabolism in Arabidopsis after the sudden onset of abiotic stresses (focusing on heat and dark). The interesting finding of our work is that KCS4, an enzyme involved in the elongation of fatty acids and wax synthesis, determines the differential accumulation of polyunsaturated triacylglycerols. By sequestering the saturated triacylglycerols into wax, KCS4 leaves free polyunsaturated triacylglycerols to accumulate in lipid bodies and to be used as an alternative source of energy.
Next steps: This approach could further be used to explore the fine-tuning of lipid metabolism in diverse environments and in other plant species.
Reference:
Urszula Luzarowska, Anne-Kathrin Ruß, Jérôme Joubès, Marguerite Batsale, Jędrzej Szymański, Venkatesh Periyakavanam Thirumalaikumar, Marcin Luzarowski, Si Wu, Feng Zhu, Niklas Endres, Sarah Khedhayir, Julia Schumacher, Weronika Jasinska, Ke Xu, Sandra Marcela Correa Cordoba, Simy Weil, Aleksandra Skirycz, Alisdair Robert Fernie, Yonghua Li-Beisson, Corina M. Fusari, Yariv Brotman. (2023). Hello darkness, my old friend: 3-KETOACYL-COENZYME A SYNTHASE4 is a branch point in the regulation of triacylglycerol synthesis in Arabidopsis thaliana https://doi.org/10.1093/plcell/koad059