Symphony of the regulators: How do plants control complex responses to environmental signals?
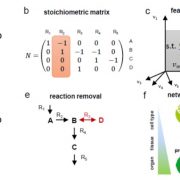
There are several models we use to conceptualize how plants respond to environmental signals through transcriptional regulation. In perhaps the best-understood model, the perception of some environmental signal flows through one or several mechanisms to a master regulator, often a transcription factor (TF). This master regulator acts as an on-off switch to trigger downstream gene expression and other plant phenotypic responses. However, some adaptive responses are not as simple as on-or-off, but instead change continuously or in a modulated response to environmental signals, as you might adjust your thermostat. Multiple regulators must be invoked to underlie these quantitative responses, suggesting a model of regulatory networks with many possible interactions among regulators. As yet we know little about how numerous, how connected, and how interactive these genetic regulatory networks are in plants.
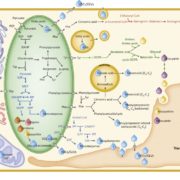
Aliphatic glucosinolates (GLS) are secondary metabolites and major contributors to plant defense against herbivores and pathogens, as well as to the distinctive mustard flavors of Brussels sprouts, horseradish, and other members of the Brassicaceae (Halkier & Gershenzon 2006). Some GLS are produced constitutively in various plant tissues, while the production of others occurs in response to developmental and environmental stimuli (Kliebenstein et al. 2002). As such, the level and composition of GLS can change dramatically with both genetic and environmental variation, and variation in GLS production can affect plant fitness, growth, and yield (Mauricio 1998; Kerwin et al. 2015). The dynamic and quantitative nature of GLS biosynthesis makes it an attractive system for understanding complex transcriptional regulation, especially given that the genes and enzymes involved are almost entirely characterized (Halkier & Gershenzon 2006).
Li et al. (2018) have taken on this challenge, focusing on epistatic (non-additive) interactions of transcriptional regulators in aliphatic GLS biosynthesis. They generated 47 double mutants of T-DNA insertion lines for 20 TFs that regulate this process. These TFs include three putative ‘master regulators’ (MYB28, MYB29, and ANT of the AP2/EREBP family) and 17 newly identified TFs of diverse gene families and annotated functions that group into six phenotypic modules based on aliphatic GLS accumulation (Li et al. 2014).
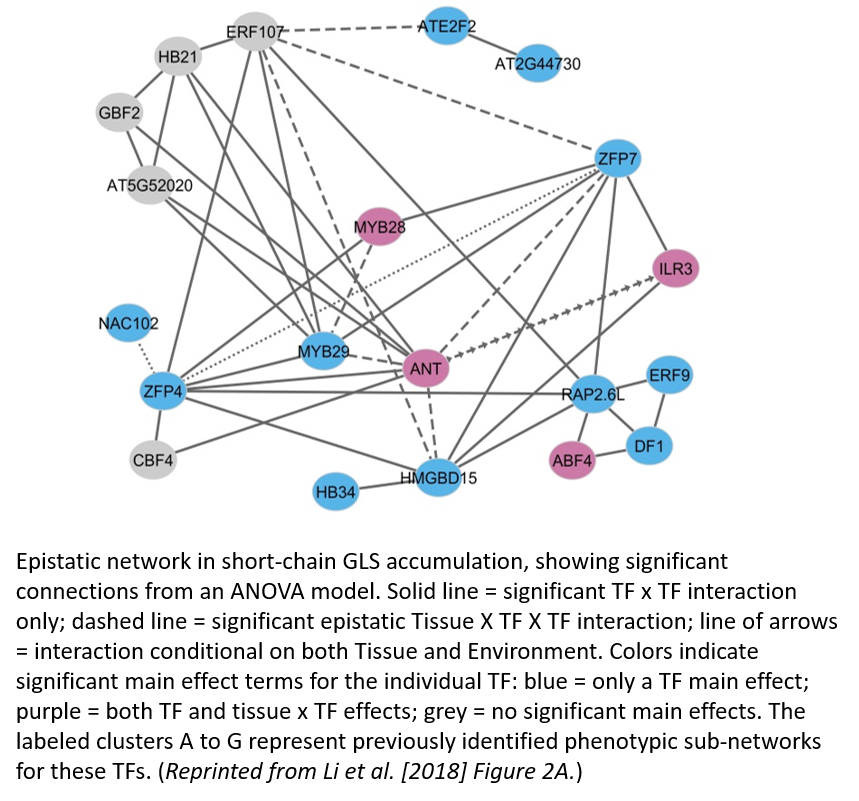 The double mutants were designed to systematically compare four types of epistasis: within phenotypic module, between phenotypic modules, interaction with MYBs, and interaction with ANT. Across these categories, in two tissues and two environments, Li et al. (2018) find widespread and largely antagonistic epistasis in the regulatory network of aliphatic GLS biosynthesis on GLS accumulation phenotypes (e.g. Figure, which shows epistasis in short chain GLS accumulation). In antagonistic epistasis, the phenotype of the double mutant is lower than the expected additive combination of each single mutant. The one exception is for the interaction of the newly identified TFs and the two MYBs, where interactions are largely additive.
The double mutants were designed to systematically compare four types of epistasis: within phenotypic module, between phenotypic modules, interaction with MYBs, and interaction with ANT. Across these categories, in two tissues and two environments, Li et al. (2018) find widespread and largely antagonistic epistasis in the regulatory network of aliphatic GLS biosynthesis on GLS accumulation phenotypes (e.g. Figure, which shows epistasis in short chain GLS accumulation). In antagonistic epistasis, the phenotype of the double mutant is lower than the expected additive combination of each single mutant. The one exception is for the interaction of the newly identified TFs and the two MYBs, where interactions are largely additive.
To examine this in more detail, Li and colleagues generated four triple mutants of myb28 myb29 (no GLS accumulation) plus one of each of four repressor TFs to look for the rescue of GLS accumulation predicted from additive effects. Instead, they observe no rescue, only the recessive epistasis expected under a master regulator model. Comparing gene expression to metabolite accumulation in mutant combinations of the three ‘master regulators’ (myb28, myb29, and ant), Li and colleagues find that epistasis in metabolite phenotype does not necessarily indicate gene expression epistasis, and vice versa.
Collectively, Li and colleagues have begun to uncover a complex and dynamic system of regulation in aliphatic GLS biosynthesis. This complexity allows for a fine-tuning of the level and composition of GLS, and suggests an orchestral model of regulation: multiple musicians (TFs) coming on and off, sometimes to dissonant or harmonious effect (epistasis), under some control by ‘conductors’ (MYBs) of the biosynthetic symphony.
References
Halkier, B. A., & Gershenzon, J. (2006). Biology and biochemistry of glucosinolates. Annu. Rev. Plant Biol. 57: 303-333.
Kerwin, R., Feusier, J., Corwin, J., Rubin, M., Lin, C., Muok, A., Larson, B., Li, B., Joseph, B., Francisco, M., Copeland, D., Weinig, C., & Kliebenstein, D. J. (2015). Natural genetic variation in Arabidopsis thaliana defense metabolism genes modulates field fitness. Elife 4: e05604.
Kliebenstein, D. J., Figuth, A., & Mitchell-Olds, T. (2002). Genetic architecture of plastic methyl jasmonate responses in Arabidopsis thaliana. Genetics 161: 1685-1696
Li, B., Gaudinier, A., Tang, M., Taylor-Teeples, M., Nham, N. T., Ghaffari, C., Benson, D. S., Steinmann, M., Gray, J. A., Brady, S. M., & Kliebenstein, D. J. (2014). Promoter-based integration in plant defense regulation. Plant Physiol. 166: 1803-1820.
Li, B., Tang, M., Nelson, A., Caligagan, H., Zhou, X., Wiest, C. C., Ngo, R., Brady, S. M., & Kliebenstein, D. J. (2018). Network-guided Discovery of Extensive Epistasis Between Transcription Factors Involved in Aliphatic Glucosinolate Biosynthesis. Plant Cell. Published January 2018. DOI: https://doi.org/10.1105/tpc.17.00805.
Mauricio, R. (1998). Costs of resistance to natural enemies in field populations of the annual plant Arabidopsis thaliana. Am. Na. 151: 20-28.