Silencing Immunity: miR159 Suppresses Pathogen Responses in Tobacco
Michael J. Skelly
To ensure optimal growth and development, plants must precisely control gene expression networks in a tissue-specific manner. MicroRNAs (miRNAs) are a class of small non-coding RNA molecules that post-transcriptionally silence genes by binding to complementary sequences in target mRNAs. Many land plant miRNA–target relationships have been strongly conserved throughout evolution due to their prominence in the regulation of fundamental developmental processes. The ability of miRNAs to diffuse across tissues allows them to act as powerful cell-to-cell messengers propagating gene silencing signals. Indeed, many developmental transcription factors are regulated by miRNAs in this manner to shape plant morphology and organ patterning (D’Ario et al., 2017).
The GAMYB gene family encodes a group of R2R3 MYB domain transcription factors that act as important transducers of gibberellin (GA) signals in both seeds and anthers (Gubler et al., 1995; Aya et al., 2009). However, in vegetative tissues, GAMYB-like genes are constitutively transcribed but strongly silenced by the miRNA miR159. Various studies in Arabidopsis using mir159 and gamyb mutants have shown that GAMYBs are not required for GA-mediated vegetative growth. However, in the absence of miR159, deregulated GAMYB expression leads to wide-ranging deleterious developmental phenotypes (Millar et al., 2019). Since the miR159-GAMYB pathway has persisted throughout evolution, this poses the question of what is the purpose of this potentially harmful constitutive GAMYB transcription?
 In this issue of Plant Physiology, Zheng et al. (2020) investigate this question by inhibiting miR159 function in both tobacco (Nicotiana tabacum) and rice (Oryza sativa). The silencing effect of miRNAs can be inhibited by ectopic expression of target mimics that sequester the endogenous miRNA. Using this approach, the authors generated transgenic plants stably expressing MIM159, an established specific target mimic for miR159 (Todesco et al., 2010; Reichel et al., 2015). Tobacco plants expressing MIM159 showed strong developmental phenotypes including stunted growth and abnormal morphologies. These effects corresponded with upregulation of GAMYB expression, confirming that the miR159-GAMYB pathway was successfully altered in MIM159 plants. The authors also generated tobacco plants overexpressing a miR159-resistant mutant GAMYB2 and found that many of these showed similar phenotypes to the MIM159 plants. Together these experiments confirm that in tobacco, miR159 also acts to prevent abnormal development by silencing GAMYB.
In this issue of Plant Physiology, Zheng et al. (2020) investigate this question by inhibiting miR159 function in both tobacco (Nicotiana tabacum) and rice (Oryza sativa). The silencing effect of miRNAs can be inhibited by ectopic expression of target mimics that sequester the endogenous miRNA. Using this approach, the authors generated transgenic plants stably expressing MIM159, an established specific target mimic for miR159 (Todesco et al., 2010; Reichel et al., 2015). Tobacco plants expressing MIM159 showed strong developmental phenotypes including stunted growth and abnormal morphologies. These effects corresponded with upregulation of GAMYB expression, confirming that the miR159-GAMYB pathway was successfully altered in MIM159 plants. The authors also generated tobacco plants overexpressing a miR159-resistant mutant GAMYB2 and found that many of these showed similar phenotypes to the MIM159 plants. Together these experiments confirm that in tobacco, miR159 also acts to prevent abnormal development by silencing GAMYB.
To establish downstream signalling components regulated by the miR159-GAMYB pathway, the authors performed RNA-Seq analysis of MIM159 tobacco plants. Strikingly, gene ontology classification showed many immune genes involved in pathogen responses were strongly upregulated in MIM159 plants compared to wild-type. Other high-ranking categories of upregulated genes included those involved in programmed cell death, and hormone and environmental stimulus. A concomitant downregulation of cell cycle-related and developmental genes was observed in MIM159 plants, likely contributing to the observed abnormal growth and morphology. Importantly, the upregulation of pathogen-responsive immune genes in MIM159 tobacco plants was associated with increased resistance to the pathogenic oomycete Phytophthora parasitica.
In the present study, Zheng et al. demonstrate that the miR159-GAMYB-mediated control of development is also conserved in rice. Similar to their results in tobacco, rice plants expressing MIM159 showed stunted growth phenotypes and elevated GAMYB expression. This suggests that at least in vegetative tissues, the requirement for miR159 suppression of GAMYB in order to ensure normal growth and development has been conserved across monocot and dicot species. However, unlike tobacco, rice MIM159 plants showed no evidence of elevated expression of pathogen-responsive immune genes. To determine if this phenomenon exists in Arabidopsis, the authors examined mir159ab plants in which two highly-expressed members of the miR159 family, miR159a and miR159b, are mutated. These plants have severe developmental defects (Allen et al, 2007), but showed no upregulation of the corresponding immune genes found in tobacco. Furthermore, miR159-resistant mutant GAMYB genes from diverse species, including tobacco, all resulted in developmental defects but no activation of immune gene expression when expressed in Arabidopsis. These results suggest that of the species tested, miR159-mediated silencing of immune responses is unique to tobacco.
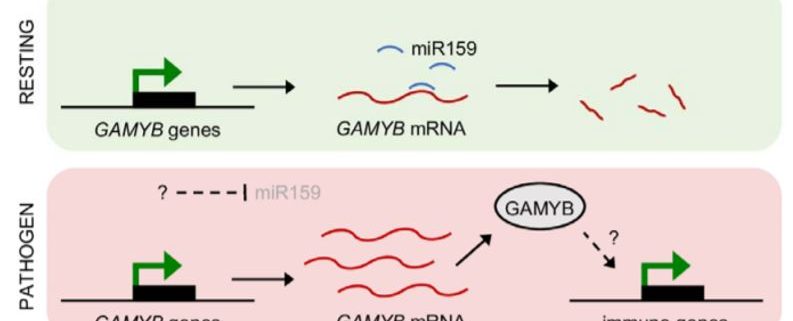
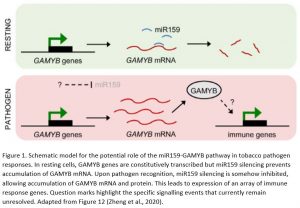
In summary, this exciting work demonstrates an unprecedented role in pathogen resistance for the miR159-GAMYB pathway in tobacco. This appears to be a fundamental role spanning different branches of plant immunity, as evidenced by the broad range of immune genes upregulated in MIM159 tobacco plants. These include genes associated with resistance against oomycetes, fungi, bacteria and viruses. Thus, the potent effects of deregulated GAMYB expression, while deleterious from a developmental standpoint, may be advantageous to tobacco when under attack from pathogens. In this model, immune responses are constitutively primed in resting cells and ready to be rapidly unleashed upon removal of miR159 silencing. Upon pathogen recognition, miR159 silencing is inhibited, triggering potent immune responses (Figure 1). However, the mechanisms by which miR159 is inhibited upon pathogen recognition and how the ensuing elevated GAMYB levels induce immune genes remain unknown. While this newly established role in immunity might explain why the miR159-GAMYB pathway is so highly conserved across the plant kingdom, evidence that this mechanism exists in species other than tobacco is scarce. Therefore, future work should aim to determine whether similar roles for miR159-GAMYB pathways exist in other plants. These efforts will likely require further identification of additional components of these complex signalling pathways and will further enhance our understanding of both miRNA-mediated gene silencing and plant immunity.
LITERATURE CITED
Allen RS, Li J, Stahle MI, Dubroue A, Gubler F, Millar AA (2007) Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc Natl Acad Sci U S A 104: 16371-16376
Aya K, Ueguchi-Tanaka M, Kondo M, Hamada K, Yano K, Nishimura M, Matsuoka M (2009) Gibberellin modulates anther development in rice via the transcriptional regulation of GAMYB. Plant Cell 21: 1453-1472
D’Ario M, Griffiths-Jones S, Kim M (2017) Small RNAs: Big Impact on Plant Development. Trends Plant Sci 22: 1056-1068
Gubler F, Kalla R, Roberts JK, Jacobsen JV (1995) Gibberellin-regulated expression of a myb gene in barley aleurone cells: evidence for Myb transactivation of a high-pI alpha-amylase gene promoter. Plant Cell 7: 1879-1891
Millar AA, Lohe A, Wong G (2019) Biology and Function of miR159 in Plants. Plants (Basel) 8
Reichel M, Li Y, Li J, Millar AA (2015) Inhibiting plant microRNA activity: molecular SPONGEs, target MIMICs and STTMs all display variable efficacies against target microRNAs. Plant Biotechnol J 13: 915-926
Todesco M, Rubio-Somoza I, Paz-Ares J, Weigel D (2010) A collection of target mimics for comprehensive analysis of microRNA function in Arabidopsis thaliana. PLoS Genet 6: e1001031