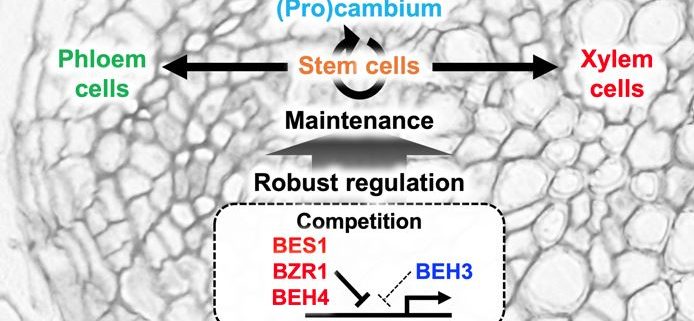
Robust control of stem cells during vascular development
Furuya et al. identify BEH3 as a stabilizer in the cambium in Arabidopsis thaliana
By Tomoyuki Furuyaa,b and Yuki Kondoa,b.
a Graduate School of Science, Kobe University, Japan
b Graduate School of Science, The University of Tokyo, Japan.
Background: Just like trees, plants like Arabidopsis undergo continuous radial growth from the activity of vascular stem cells in the cambium. Although many efforts have been poured into uncovering vascular stem cell regulators by genetics and transcriptomics methods, the molecular mechanism enabling the robust regulation of vascular stem cells was not fully understood. We previously established an ectopic vascular cell differentiation system, VISUAL, which efficiently mimics the entire process of vascular development by the addition of phytohormone and chemical compounds. Through genetic analyses using VISUAL, we identified the brassinosteroid-related transcription factors BES1 and BZR1 as major regulators of vascular stem cell maintenance.
Question: We wished to discover novel regulators of vascular stem cell maintenance to understand how vascular stem cells can be maintained locally with such a robust control, while allowing cell differentiation further away.
Findings: By combining multiple transcriptome datasets of the progression of vascular cell differentiation derived from VISUAL, we identified a new set of 394 cambium-related genes based on a gene co-expression network. Of those, we demonstrate here that the BES/BZR homolog BEH3 is a potential stem cell regulator. beh3 mutants exhibited a larger variation in vascular size than the wild type, suggesting a role for BEH3 as a stabilizer of vascular development. BEH3 also functioned antagonistically with BES1 in vascular development. At the biochemical level, BEH3 had a weaker transcriptional repressor activity than other BES/BZR homologs. Nevertheless, BEH3 competed with them to bind to the same DNA motif, thereby passively interfering with the activity of other members. Mathematical modeling revealed that the competitive relationship between BES/BZR members contributes to minimizing “phenotypic variation” in vascular size.
Next steps: Our co-expression network analysis can uncover informative gene lists to examine whether genes of interest are expressed in xylem, cambium or phloem tissues. Moreover, we identified BEH3 as a non-canonical member in the BES/BZR family. Further comparative analyses will help uncover the mechanism(s) underlying the functional differences between BEH3 and other BES/BZR homologs.
Citation: Tomoyuki Furuya, Masato Saito, Haruka Uchimura, Akiko Satake, Shohei Nosaki, Takuya Miyakawa, Shunji Shimadzu, Wataru Yamori, Masaru Tanokura, Hiroo Fukuda, Yuki Kondo (2021). Gene co-expression network analysis identifies BEH3 as a stabilizer of secondary vascular development in Arabidopsis. Plant Cell, https://doi.org/10.1093/plcell/koab151