Review: Copy Number Variations shaping plant domestication (Trends Plant Sci)
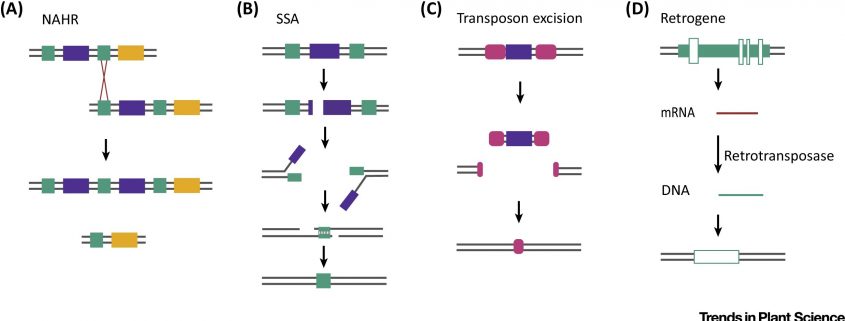
Human-associated plant domestication is a co-evolutionary process that began at least 12,000 years ago. However, the genetic variations underlying many domestication traits are still unknown. In this review, Lye and Purugganan discuss how copy number variations (CNVs) have played an important role in plant and animal domestication. CNVs are deletions, duplications or amplifications (usually at least 1 kb long) that can occur in both coding and noncoding genomic regions. Genetic mechanisms such as unequal cross over, DNA double-strand break repair and transposon activity – all of these can give rise to CNVs. While deletion CNVs are often associated with domestication traits, gene amplifications are more common in CNVs linked to diversification traits and environmental adaptation. Understanding the role of CNVs in the evolution of domestication traits can thus provide useful resources for crop improvement. (Summary by Saima Shahid) Trends in Plant Sci. 10.1016/j.tplants.2019.01.003.