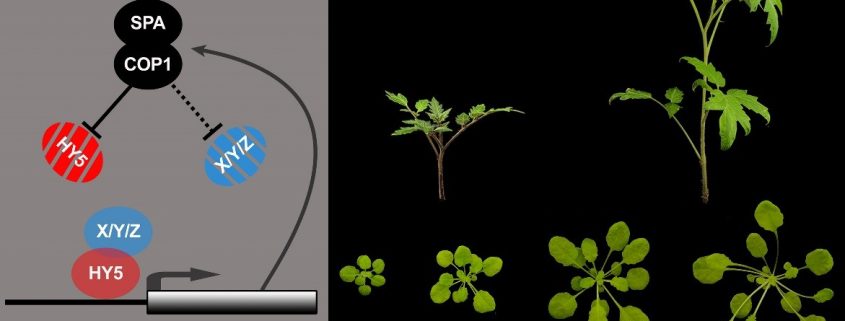
HY5 Activates Rather Than Represses its Direct Targets, Including its Negative Regulator SPA1
Burko et al. carry out detailed phenotypic and molecular analyses of constitutive activator and repressor HY5 fusion proteins to identify its direct targets. Plant Cell https://doi.org/10.1105/tpc.19.00772
Background: Plants are highly responsive to environmental factors, and differences in light, temperature, nutrients, or other conditions can change how they grow. To form their final shape, plants collect all of this information and use many proteins called Transcription Factors (TFs) to control the expression of hundreds and sometimes thousands of genes. One such TF, called HY5, is a central player in the plant’s response to light. HY5 regulates cell elongation, pigment accumulation, flowering, root development, and many other processes. Plant biologists think HY5 does this by binding to many genes in the genome and turning them on or off. But the exact mechanism of HY5 activity is still unknown, and some of the explanations for how the process works don’t agree.
Question: TF activity can be difficult to define, and earlier studies were not able to clearly show how HY5 works. We made new versions of HY5 that could only turn on or only turn off gene expression and asked if these tools could help us understand how HY5 does its job.
Findings: First, we tested these new HY5 variants in the model plant Arabidopsis thaliana. We compared many aspects of growth, gene expression, and DNA binding in these plants. Using these data, we generated a list of high confidence HY5 direct targets including members of the COP1/SPA complexes which repress HY5 activity, revealing a light-regulated feedback mechanism. We provide data supporting a HY5-SMZ/SNZ-flowering control pathway, potentially explaining why hy5 mutants flower early. We also show that multiple properties of the HY5-COP/SPA feedback loop can be used to control the growth of tomato plants. Additionally, we show that HY5 works to turn on gene expression, in contrast to the prevailing model of both on and off activity. This enables more detailed analysis of the HY5-dependent gene expression networks.
Next steps: More detailed follow-up on the new HY5 direct targets is needed. Understanding how the HY5/COP1-SPA feedback loop can be tuned to optimize plant size also requires more study in tomato and other crops. To do this, we need to make sure HY5 is expressed in the right part of the plant at the right time.
Burko, Y., Seluzicki, A., Zander, M., Pedmale, U., Ecker, J.R., and Chory, J. (2020). Chimeric activators and repressors define HY5 activity and reveal a light-regulated feedback mechanism. Plant Cell https://doi.org/10.1105/tpc.19.00772