Dynamic 3D genome architecture during polyploidy and domestication
By Longfei Wang and Qingxin Song
Background: Soybean and common bean are important sources of vegetable protein for humans and livestock. These two legumes originated from a common ancestor that underwent a whole-genome duplication (WGD) event ~56.5 million year ago (MYA). After diverging from common bean ~19.2 MYA, soybean experienced an independent WGD event at ~10 MYA and was domesticated in east Asia 6000–9000 years ago. Domestication resulted in higher seed yield and wider geographical distribution, accompanied by extraordinary morphological changes including loss of pod shattering, erect and compact stem architecture, and reduction in photoperiod sensitivity. Multiple dimensions of genome organization play critical roles in animal and plant development; however, we know relatively little about the dynamic patterns of chromatin architecture during polyploidy and crop domestication.
Question: We wanted to know how chromatin architecture changes during post-polyploidy diploidization (soybean vs common bean) and soybean domestication (cultivated soybean vs wild soybean). We also hoped to reveal the relationships between chromatin interaction, epigenetic marks, chromatin accessibility, and transcript abundance during evolution and domestication of paleopolyploid soybean.
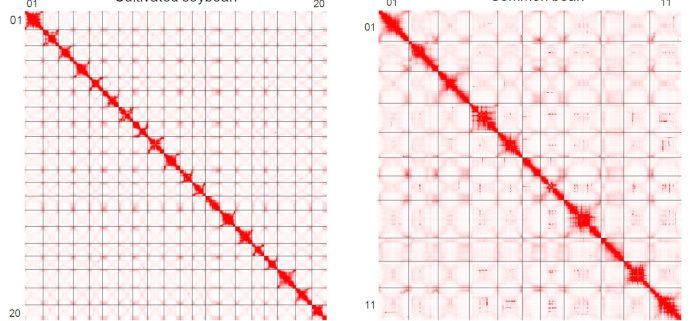
Findings: We performed in situ Hi-C experiments using young leaves to uncover the 3D chromatin structures of common bean, wild soybean, and cultivated soybean. We found chromatin architecture showed dramatic changes during polyploidization and subsequent diploidization. Compared with single-copy and small-scale duplicated genes, WGD genes displayed more long-range chromosomal interactions and were coupled with higher levels of gene expression and chromatin accessibilities. Chromatin loop reorganization occurred in many genes during soybean domestication. Interestingly, these genes tended to show different expression levels between wild and cultivated soybean. Moreover, genes with chromatin loops were under stronger artificial selection than genes without loops in soybean.
Next steps: We plan to investigate convergence and divergence of chromatin architecture among soybean varieties. Furthermore, we want to study the role of chromatin architecture in response to various environmental stresses.
Longfei Wang, Guanghong Jia, Xinyu Jiang, Shuai Cao, Z. Jeffrey Chen and Qingxin Song (2021) Altered chromatin architecture and gene expression during polyploidization and domestication of soybean. Plant Cell. https://doi.org/10.1093/plcell/koab081