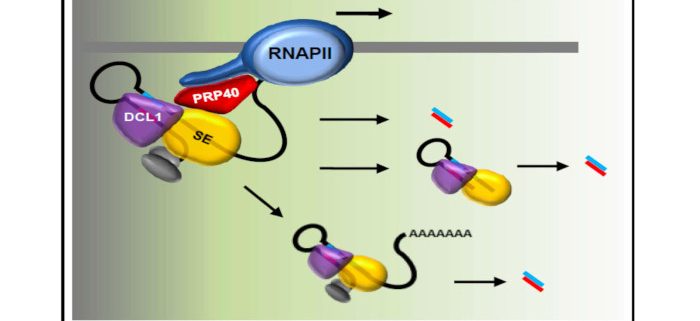
Co-transcriptional processing of microRNA precursors
Stepien et al. explore how an auxiliary U1 snRNP protein interacts with RNA polymerase II to regulate co-transcriptional processing of microRNAs. https://doi.org/10.1093/plcell/koac278
Background: MicroRNAs (miRNAs) are single-stranded, usually 21-nt-long, RNAs that regulate basic developmental processes as well as plant responses to fluctuations in environmental conditions. Plant miRNA genes (MIRs) are transcribed by RNA polymerase II (RNAPII) to generate long precursors that are processed into mature miRNAs by the miRNA biogenesis complex known as the microprocessor.
Question: We have previously demonstrated that processing of plant miRNA precursors takes place already during their synthesis by RNAPII. However, the mechanism of regulation of microprocessor assembly on primary transcripts of MIRs was unknown.
Findings: In these studies, we show that the auxiliary U1 snRNP protein AtPRP40 interacts directly with the C-terminal domain (CTD) of RNAPII as well as one of the microprocessor components, and coordinates the binding of microprocessor elements to miRNA primary precursors at the sites of their synthesis. We show that the impaired cotranscriptional microprocessor assembly that is observed in the absence of AtPRP40 is accompanied by RNAPII accumulation at miRNA genes and retention of miRNA precursors at their transcription sites. Our data suggest that cotranscriptional microprocessor assembly, regulated by AtPRP40, positively affects RNAPII transcription of miRNA genes.
Next steps: The obtained results raise interesting questions about a role of AtPRP40 in coordinating transcription and other processing events. Further studies will explore the influence of AtPRP40 on processes other than miRNA biogenesis.
Reference:
Agata Stepien, Jakub Dolata, Tomasz Gulanicz, Dawid Bielewicz, Mateusz Bajczyk, Dariusz J. Smolinski, Zofia Szweykowska-Kulinska, Artur Jarmolowski (2022) Chromatin-associated microprocessor assembly is regulated by the U1 snRNP auxiliary protein PRP40 https://doi.org/10.1093/plcell/koac278