An E3 ubiquitin ligase modulates brassinosteroid signaling by targeting the transcription factor BES1 to selective autophagy under sucrose starvation
Wang et al. identify a mechanism by which plants reduce growth under sucrose starvation conditions. Plant Cell. https://doi.org/10.1093/plcell/koab210
by Ping Wang and Yanhai Yin (Iowa State University)
Background: Plants fine-tune their growth and stress-response programs to adapt to the environment. Brassinosteroids (BRs), a major family of steroid growth-promoting plant hormones, are implicated in stress responses. BR regulates the protein level and activity of the transcription factor BRI1-EMS-SUPPRESSOR 1 (BES1), which is crucial for plant growth and response to stresses or suboptimal growth conditions. Another crucial response to stress conditions is autophagy (‘self-eating’), which functions in the degradation and recycling of proteins or organelles in the cells to recycle nutrients during stress. For example, insufficient light causes stress and potentially sucrose starvation; plants respond to low light by slowing their growth and inducing autophagy. We previously found that BES1 was targeted to selective autophagy by the ubiquitin receptor DSK2 upon sucrose starvation to slow down plant growth.
Question: We wanted to know how BES1 is marked by ubiquitin that targets proteins to degradation, what is the E3 ubiquitin ligase that marks BES1 for recycling, how does this modification lead to selective autophagy of BES1, and whether or not this process is dependent on DSK2 under sucrose starvation.
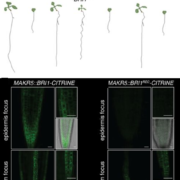
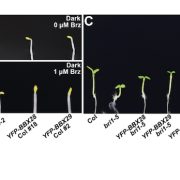
Findings: We identified a previously uncharacterized F-box family E3 ubiquitin ligase called BES1-ASSOCIATED F-BOX1 (BAF1). BAF1 interacts with BES1 and mediates its ubiquitination and degradation. BAF1-mediated BES1 degradation through selective autophagy requires DSK2-dependent. Overexpression of BAF1 in Arabidopsis increased tolerance to sucrose starvation but somewhat reduced plant growth, while the loss-of-function baf1 mutant displayed the opposite phenotypes (see image).
Next steps: BAF1 targets BES1 to selective autophagy under sucrose starvation, but whether or not BAF1 is regulated by the stress remains to be determined. Our work not only established BAF1 as an E3 ubiquitin ligase, but also revealed a mechanism for plants to reduce growth during suboptimal conditions. It will be interesting to explore whether BAF1 can be manipulated to achieve optimal crop production under stress conditions.
Reference:
Ping Wang, Trevor M Nolan, Natalie M Clark, Hao Jiang, Christian Montes-Serey, Hongqing Guo, Diane C Bassham, Justin W Walley, Yanhai Yin (2021) The F-box E3 ubiquitin ligase BAF1 mediates the degradation of the brassinosteroid-activated transcription factor BES1 through selective autophagy in Arabidopsis. Plant Cell https://doi.org/10.1093/plcell/koab210