What happens when DNA breaks are repaired and when repair fails?
Samach et al. use a pigment-based visual assay to explore the repair of double-stranded DNA breakd in plants.
https://doi.org/10.1093/plcell/koad209
Avraham Levy and Aviva Samach, Weizmann Institute of Science, Rehovot, Israel.
Background: Plant breeding is the art of combining desirable traits from parental genomes. This normally occurs during meiosis through crossover, the exchange of chromosomal segments between homologous chromosomes. So far it has not been possible to target the precise breakpoints of crossovers at meiosis. Somatic crossover is rare, but we previously showed that it can be stimulated by a DNA double-strand break (DSB) and can be transmitted to the next generation. This was shown in tomatoes (Solanum lycopersicum) at specific fruit-color loci, but a more general system is needed to become a useful tool for precise breeding.
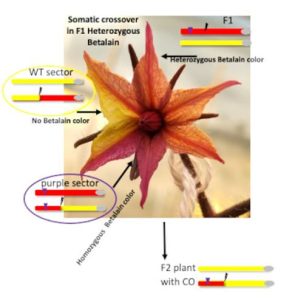
Question: To better understand somatic DSB repair via homologous recombination, we developed an assay enabling visualization of somatic crossover via segregation of a transgenic purple color marker (the Betalain pigment), at a broad range of loci. Through this process, heterozygous tissues become homozygous, forming wild-type or dark purple sectors in tomato leaves, flowers or fruits.
 Findings: We confirmed that somatic crossover can be detected visually at a CRISPR-mediated DSB site, opening the prospect for precise breeding in crops. Moreover, we showed that loss of heterozygosity could be due to major chromosomal rearrangements triggered by defective repair of the DSB. Plants where this occurred contained large deletions and translocations and were sterile, showing micronuclei as well as bridges in dicentric chromosomes, at meiosis, as described in McClintock’s Breakage-Fusion-Bridge Cycle triggered by transposons. This genomic reshuffling is similar to chromothripsis in mammalian cells. Both crossover and chromothripsis events were rare but could be detected thanks to the visual assay.
Findings: We confirmed that somatic crossover can be detected visually at a CRISPR-mediated DSB site, opening the prospect for precise breeding in crops. Moreover, we showed that loss of heterozygosity could be due to major chromosomal rearrangements triggered by defective repair of the DSB. Plants where this occurred contained large deletions and translocations and were sterile, showing micronuclei as well as bridges in dicentric chromosomes, at meiosis, as described in McClintock’s Breakage-Fusion-Bridge Cycle triggered by transposons. This genomic reshuffling is similar to chromothripsis in mammalian cells. Both crossover and chromothripsis events were rare but could be detected thanks to the visual assay.
Next steps: It will be important to better understand what determines the fate of a DSB; when repair leads to crossover or alternatively, to chromothripsis. The new assay will also enable to study how targeted somatic crossover can become more efficient for breeding applications.
Reference:
Aviva Samach, Fabrizio Mafessoni, Or Gross, Cathy Melamed-Bessudo, Shdema Filler-Hayut, Tal Dahan-Meir, Ziva Amsellem, Wojciech P. Pawlowski, and Avraham A. Levy (2023) CRISPR/Cas9-induced DNA breaks trigger crossover, chromosomal loss, and chromothripsis-like rearrangements. https://doi.org/10.1093/plcell/koad209



