Substrate recognition by asparaginyl peptide ligases becomes crystal clear
By Side Hu, Abbas El Sahili and Julien Lescar
Background: Proteases, protein scissors that cut proteins into smaller pieces, can be found almost everywhere in the natural world, including in plants. However, protein ligases, which stitch together small pieces of proteins together, are much harder to find. Yet protein ligases such as peptide asparaginyl ligases (PALs) are very valuable, because they have many applications in biomedical research, such as conjugating proteins with a label for imaging, or fusing an antibody with a toxin to specifically kill cancer cells in patients.
Question: How do PALs recognize their peptide substrates? We studied VyPAL2, a PAL from the medicinal plant Viola yedoensis with very strong ligase activity at neutral pH. We also asked how VyPAL2 can cut itself into two pieces at acidic pH, so that it becomes active.
Findings: We expressed the VyPAL2 protein in insect cells and obtained crystals of the active protein. Comparing our crystal structures with published protease structures, we found a backbone shift induced by a specific residue that acts as an important activity determinant. We also found that the catalytic cysteine, which was reported to be the key residue for activation, is not necessary for maturation. Instead, a catalytic histidine plays a key role and we came up with a scheme to account for auto-activation.
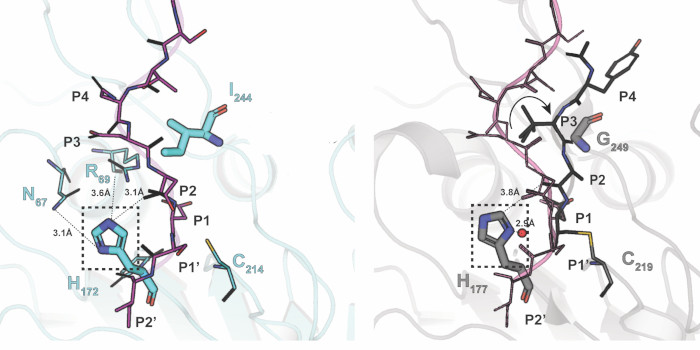
The asparaginyl peptide ligase VyPAL2 (green ribbon bottom) is captured in the act of binding a peptide substrate (sticks with residues labeled) projecting from another VyPAL2 molecule (red ribbons).
Next steps: The results described here will help engineer ligases with altered specificity that can be used in biotechnology and precision medicine.
Reference:
Side Hu, Abbas El Sahili, Srujana Kishore, Yee Hwa Wong, Xinya Hemu, Boon Chong Goh, Zhipei Sang, Zhen Wang, James P Tam, Chuan-Fa Liu, Julien Lescar (2022) Structural basis for proenzyme maturation, substrate recognition, and ligation by a hyperactive asparaginyl peptide ligase



