Interdependence of subunits of the m6A methyltransferase complex
By Lisha Shen, Temasek Life Sciences Laboratory and Department of Biological Sciences, National University of Singapore, Singapore
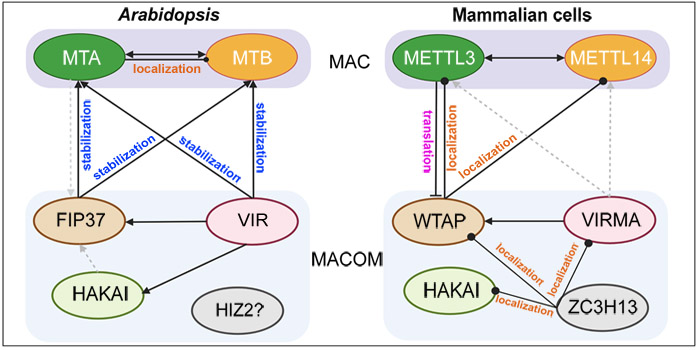
Background: N6-methyladenosine (m6A), methylation at the nitrogen-6 position of adenosine, is an abundant internal modification of messenger RNA (mRNA) in plants, animals, and fungi. Adding m6A to mRNAs requires an evolutionarily conserved multicomponent complex called the m6A methyltransferase complex. In the model plant Arabidopsis thaliana, this complex contains two core methyltransferases, mRNA adenosine methylase (MTA) and MTB, and several accessory subunits including FKBP12 INTERACTING PROTEIN 37KD (FIP37), VIRILIZER (VIR) and HAKAI. Among these components, MTA, MTB, FIP37, and VIR are essential for m6A methylation.
Question: Although several components have been identified in the m6A methyltransferase complex, how these components influence each other to fulfill the function of the m6A methyltransferase complex as a whole remains largely unknown.
Findings: The individual components of the m6A methyltransferase complex mostly affect each other at the protein level rather than at the gene expression level. First, MTA and MTB affect each other’s protein abundance. Second, FIP37 and VIR stabilize MTA and MTB, thus maintaining m6A methyltransferase complex function. Third, VIR affects the protein accumulation of FIP37 and HAKAI. In contrast, HAKAI has little effect on the protein abundance of MTA, MTB and FIP37. These results suggest that modulation of protein levels of individual subunits at the post-translational level is essential for the proper function of the m6A methyltransferase complex in depositing m6A modifications in plants.
Next steps: In future studies, I will further investigate whether protein homeostasis among various subunits of the m6A methyltransferase complex is dynamically altered and regulated in different developmental contexts and in response to different environmental conditions.
Reference:
Lisha Shen. (2023). Functional interdependence of subunits of the m6A methyltransferase complex in Arabidopsis. https://doi.org/10.1093/plcell/koad070