Autophagy: Both Friend and Foe in Pseudomonas Infection
Eukaryotes use two major systems for getting rid of unwanted proteins: the ubiquitin proteasome system and autophagy. The proteasome degrades ubiquitinated proteins. Autophagosomal vesicles encapsulate cellular waste and either deliver it to the vacuole or fuse with a lysosome. Animal cells use autophagy for pathogen defense; however, some pathogens have evolved to exploit autophagy to increase their own pathogenicity (Mostowy, 2013). In plants, much less is known about the role of autophagy in infection.
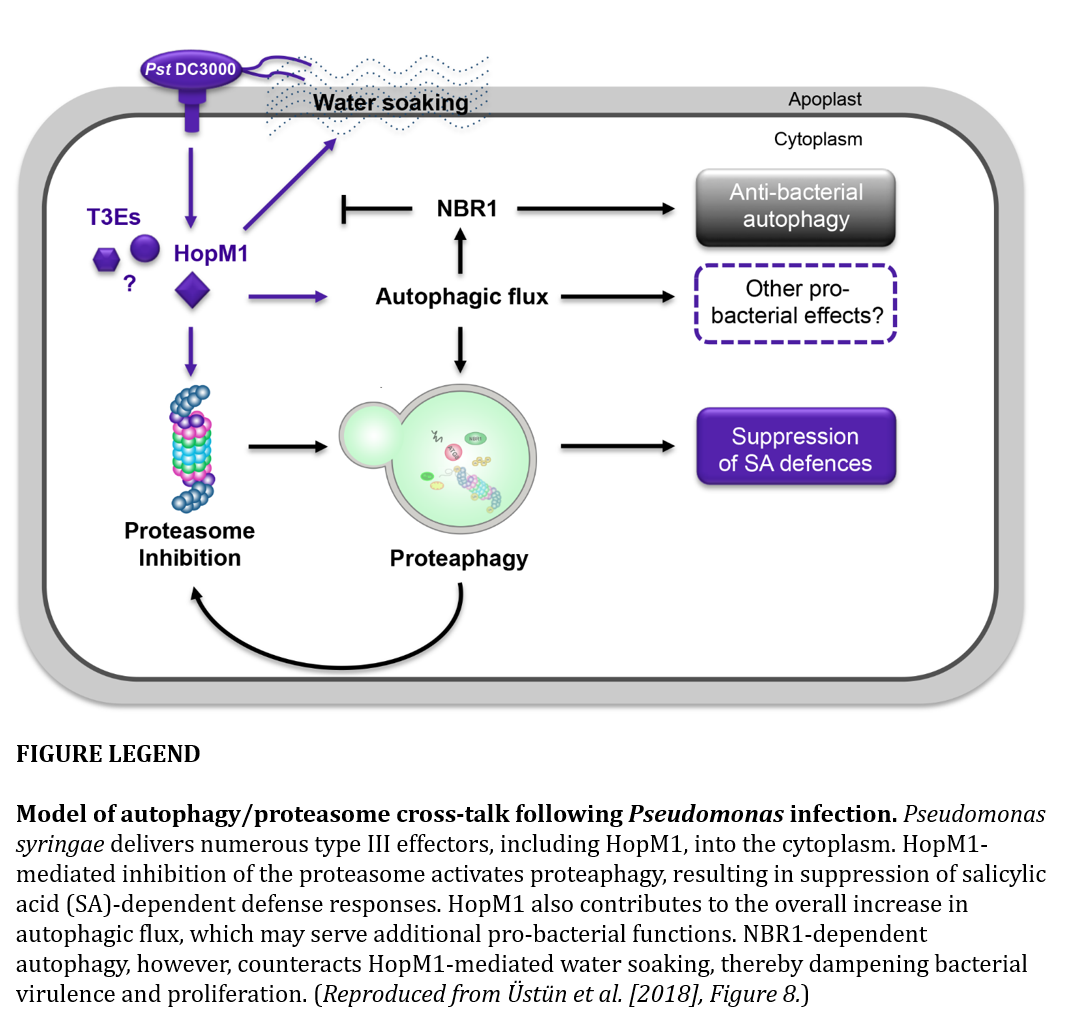
Üstün et al. (2018) show that when virulent Pseudomonas syringae infects Arabidopsis, two main things happen with respect to autophagy: 1) The bacteria activates autophagy via the type-III effector HopM1 to remove proteasomes enabling increased proliferation. 2) An NBR1-dependent autophagic response in the plant decreases disease progression. Autophagy thus plays a role in both the bacteria’s pathogenesis and the host’s immune response, yet through different selective autophagy pathways.
The authors first showed that infection by P. syringae pv. tomato DC3000 suppressed proteasome activity in wild-type Arabidopsis plants. Conversely, infection led to an increase in proteasome activity in mutant plants missing core autophagy machinery. This suggests that Pseudomonas infection causes proteasome suppression via an autophagy mechanism.
Key mutations were used to further demonstrate proteasome and autophagy crosstalk. Autophagy mutants were more resistant to Pseudomonas infection unless crucial subunits of the proteasome were mutated. Ubiquitinated proteins also accumulated in autophagy mutants. Further, infected plants lacking a key proteasome-specific autophagy receptor did not exhibit proteasome suppression.
Several lines of evidence presented suggest that Pseudomonas infection activates autophagy to suppress the proteasome and increase pathogenesis. Transcription of a core autophagy gene (ATG8) and the autophagy receptor NBR1 was increased following Pseudomonas infection. Several genes that are known to be upregulated in response to proteasome inhibition were also upregulated in Pseudomonas-infected plants. A cell biology approach was used to confirm that autophagy proteins associate with the proteasome following infection.
The authors suggest that the effect of Pseudomonas infection on autophagy occurs via type-III effector proteins. A strain of Pseudomonas lacking a type-III secretion system had different effects on NBR1 and ATG8 than did wild-type bacteria. Expression of HopM1, an effector known to inhibit proteasome capacity (Üstün et al., 2016), lead to increased autophagic flux. HopM1 also colocalized with proteasome subunit aggregations and with ATG8 and NBR1. Finally, a Pseudomonas strain lacking 28 effectors, but with HopM1 reintroduced, behaved similarly to wild-type. Transient expression of HopM1 in plants also lead to Pseudomonas infection-like symptoms. Together, these results suggest that the type-III effector HopM1 is responsible for autophagy-induced proteasome suppression.
To test the biological relevance of NBR1 over-accumulation following infection, Üstün et al. challenged nbr1-2 mutants with Pseudomonas. The mutant plants were far more sensitive to infection; however, transient over-expression of NBR1 led to a decrease in lesions. This effect was dependent on the ubiquitin-binding domain and ATG8-interacting motif of NBR1, suggesting that NBR1 contributes to autophagic degradation of one or more factors important for lesion formation. HopM1 levels were unchanged by NBR1, suggesting that it targets some other unknown factor(s). The authors conclude that NBR1 selective autophagy acts independently of the pathogen-induced proteaphagy mechanism and instead contributes to host defense.
The long evolutionary history of plants and bacteria pathogens has lead to an arms race where a plant’s own defenses are sometimes used against it. This appears to be the case with Pseudomonas and autophagy. While the plant uses specific autophagic degradation to prevent the formation of disease-promoting “water-soaked” lesions, the bacteria hijacks the system to suppress the proteasome.
REFERENCES
Mostowy, S. (2013). Autophagy and bacterial clearance: a not so clear picture. Cell Microbiol 15, 395-402.
Üstün, S., Hafrén, A., Liu, Q., Marshall, R.S., Minina, E.A., Bozhkov, P., Vierstra, R.D., and Hofius, D. (2018). Bacteria exploit autophagy for proteasome degradation and enhanced virulence in plants. Plant Cell. https://doi.org/10.1105/tpc.17.00815.
Üstün, S., Sheikh, A., Gimenez-Ibanez, S., Jones, A., Ntoukakis, V., and Börnke, F. (2016). The Proteasome Acts as a Hub for Plant Immunity and Is Targeted by Pseudomonas Type III Effectors. Plant Physiol 172, 1941-1958.



