A Root Endoploidy Map: The Spatial and Temporal Arrangement of Dividing and Endocycling Cells
Bhosale et al. computationally predicted and experimentally verified the DNA ploidy level of all cells in the Arabidopsis root tip, revealing that endoreplication is spatiotemporally regulated, stress-responsive, and likely important to coordinate cell expansion with changes in cell wall structure. Plant Cell https://doi.org/10.1105/tpc.17.00983
By Rahul Bhosale, Steven Maere and Lieven De Veylder
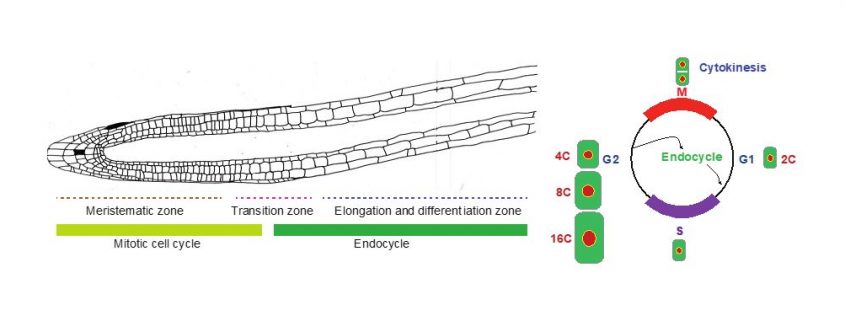
Background: Every new cell arises from division of an existing cell. In plants, these cell divisions mainly occur at the tip of shoots and roots. After a few rounds of cell division, cells stop dividing and differentiate, while simultaneously increasing their size. In many plant species, the start of differentiation is accompanied by the onset of endoreplication, an alternate cell cycle during which cells duplicate their genome without cell division, resulting in a duplication of their DNA content, i.e., an increase in ploidy level. In the root tip of the model plant species Arabidopsis thaliana, up to three endocycles can be observed, resulting in cells with a 2C, 4C, 8C, or 16C DNA content. The biological significance of these endoploidy increases is still under debate.
Question: Although we know that many plant species undergo endoreplication, our knowledge of which cell types become polyploid and how cells with different endoploidy levels are integrated into a developing organ is limited. Current protocols that quantify DNA ploidy are either time-consuming or lack tissue resolution. We addressed this problem by combining biological data with computational modeling.
Findings: Through the integration of different experimental data sets into a mathematical model, we pinpointed a set of 332 marker genes that reliably predict a three-dimensional DNA ploidy map for the complete root tip of Arabidopsis. The map obtained revealed that the level of endoreplication is to a remarkable extent differentially controlled across the distinct root tissues and their developmental stages. Moreover, a simplified version of the mathematical model allowed us to predict the effect of 149 different stress treatments on the root’s endoploidy distribution, and our results suggest that the root’s DNA ploidy pattern is highly stress-responsive. Furthermore, we demonstrated that endoreplication triggers the expression of cell wall-modifying genes, which might function to safeguard the cell’s structural stability upon turgor-driven cell expansion.
Next steps: We are currently aiming to understand exactly how the spatiotemporal root endoreplication pattern changes under various stresses and whether these changes are linked to stress adaptation. We are also studying how the endoploidy level of a cell controls expression of cell wall biosynthesis and cell wall-modifying genes.
Rahul Bhosale, Veronique Boudolf, Fabiola Cuevas, Ran Lu, Thomas Eekhout, Zhubing Hu, Gert Van Isterdael, Georgina M. Lambert, Fan Xu, Moritz K. Nowack, Richard S. Smith, Ilse Vercauteren, Riet De Rycke, Veronique Storme, Tom Beeckman, John C. Larkin, Anna Kremer, Herman Höfte, David W. Galbraith, Robert P. Kumpf, Steven Maere, Lieven De Veylder. (2018). A Spatiotemporal DNA Endoploidy Map of the Arabidopsis Root Reveals Roles for the Endocycle in Root Development and Stress Adaptation. Plant Cell 30: 2330-2351; DOI: https://doi.org/10.1105/tpc.17.00983.