Regulation and Function of a Plant Energy Sensor
Ramon et al. show how plants have modified the ancient, highly conserved eukaryotic energy sensor to better fit their unique lifestyle and cope with changing environmental conditions. Plant Cell https://doi.org/10.1105/tpc.18.00500
By Nathalie Crepin and Filip Rolland, KU Leuven, Belgium
Background: All living organisms need to maintain a positive energy balance for survival and growth. This is especially challenging for plants, which are both immobile and self-supporting (through photosynthesis) and are typically exposed to continuously changing environmental conditions and diverse types of stress. Eukaryotic organisms (including animals, fungi, and plants) use a specific type of protein kinase as a cellular fuel sensor, called SnRK1 in plants. These kinases activate energy-producing (catabolic) reactions and repress energy-consuming (anabolic) processes when carbon and energy supplies become limited. They typically operate as complexes with a catalytic α subunit (the actual kinase) and regulatory b and g subunits.
Question: While the overall structure and function of these proteins are largely conserved, the diverse lifestyles of different eukaryotic organisms require different regulatory mechanisms. We used cellular assays and mutant and transgenic plants to investigate exactly how the activity of the plant SnRK1 kinase is regulated.
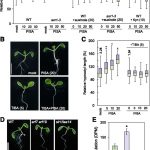
Findings: Unlike its yeast and animal counterparts, the plant SnRK1 kinase has regulatory subunit-independent and auto-phosphorylation (auto-activation) activity. This indicates that plants activate their metabolic stress response by default and that this response is repressed when sufficient carbon and energy are available. Such negative regulation (with de-repression rather than activation) is more generally used by plants and is possibly a faster and/or more reliable strategy to cope with rapidly changing environmental conditions. Low energy stress (and thus de-repression) causes translocation of the SnRK1 kinase subunit to the nucleus to induce target gene expression. The membrane-associated regulatory b subunits can restrict this nuclear localization. Analysis of transgenic plants with altered SnRK1 localization confirmed the importance of this regulatory mechanism to replenish cellular energy for survival but also revealed important functions for the plant kinase in normal growth and development.
Next steps: Future research will focus on further elucidating the upstream regulatory mechanisms (SnRK1 complex dynamics and subcellular localization) and the downstream pathways responsible for the regulation of growth and development by nuclear and cytoplasmic SnRK1. This knowledge may contribute to increasing crop yield by uncoupling stress tolerance from the typically associated (energy-saving) reduction in growth.
Matthew Ramon, Tuong Vi T. Dang, Tom Broeckx, Sander Hulsmans, Nathalie Crepin, Jen Sheen, and Filip Rolland (2019). Default Activation and Nuclear Translocation of the Plant Cellular Energy Sensor SnRK1 Regulate Metabolic Stress Responses and Development. Plant Cell 31: xx; https://doi.org/10.1105/tpc.18.00500
Key words: SnRK1, energy sensor, metabolic stress