Pathogenic Bacteria Use A “Self-Eating” Process to Trick Plants
Üstün et al. show that pathogenic Pseudomonas enhances the autophagic turnover of proteins in Arabidopsis https://doi.org/10.1105/tpc.17.00815.
By Suayib Üstün and Daniel Hofius
Background: Autophagy and the proteasome are the major pathways for protein degradation in eukaryotes. While the proteasome is a very specific degradation system, autophagy has been considered as an unspecific “self-eating” process to recycle cell content. However, evidence has accumulated that autophagy can also act as a selective pathway that engages autophagy receptors (such as NBR1) to sequester specific cargo (proteins) into specialized vesicles, so-called autophagsomes, for subsequent delivery into digestive organelles (such as the vacuole in plant cells). Both degradation pathways control many cellular processes during development and in response to environmental stimuli, including pathogens. In plants, the proteasome is essential for defense against bacteria, and thus is targeted and inhibited in its function by bacterial proteins (so-called type-III effectors) to promote disease. By contrast, the role of autophagy in plant-bacteria interactions is not well understood.
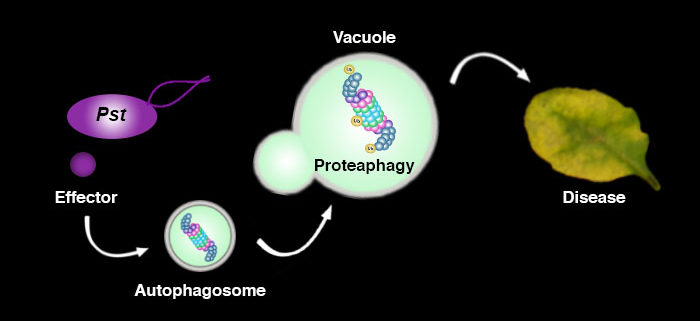
Question: Previous studies suggested that defective proteasomes are recycled by autophagy in a process termed proteaphagy. Based on the knowledge that bacteria are able to suppress proteasome function, we hypothesized that bacteria “hijack” the proteaphagy pathway to enhance virulence and disease.
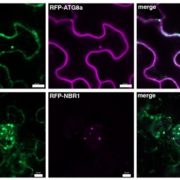
Findings: Our study demonstrates that the plant bacterium Pseudomonas syringae pv. tomato DC3000 (Pst) activates autophagy and enhances the autophagic turnover of proteins (i.e., autophagic flux) in the model plant Arabidopsis thaliana for the benefit of infection. This effect was dependent on the delivery of bacterial type-III effectors inside the plant cell. We show that that Pst effector HopM1 stimulates autophagy and the removal of proteasomes. In contrast to the pro-bacterial effects of proteaphagy, the NBR1-mediated selective autophagy pathway activated during bacterial infection counteracts disease progression, as illustrated by the restriction of HopM1-induced water-soaked lesions. We propose that distinct selective autophagy pathways with pro- and anti-bacterial functions operate in response to bacterial attack and determine the infection outcome.
Next steps: Given the established role of enhanced autophagic turnover during bacterial infection, we have indications that besides the proteasome other proteins are degraded by Pst-induced autophagy. One major task for the future is to identify new autophagy cargo proteins to more fully understand the nature of bacterial disease and plant resistance.
Suayib Üstün, Anders Hafrén, Qinsong Liu, Richard S. Marshall, Elena A. Minina, Peter V. Bozhkov, Richard D. Vierstra, and Daniel Hofius (2018). Bacteria Exploit Autophagy for Proteasome Degradation and Enhanced Virulence in Plants. Plant Cell 30: 668-685; DOI: https://doi.org/10.1105/tpc.17.00815