Plants on (Brassinos)steroids: Degradation of the Transcription Factor BZR1 by the E3 Ubiquitin Ligase PUB40
Brassinosteroids (BR) are the only plant hormones with an animal counterpart: steroid hormones. Analysis of BR signaling demonstrated that steroid receptors could reside at the plasma membrane and modulate growth and gene expression, an observation that is now becoming appreciated in non-plant systems (Levin and Hammes, 2016). Our understanding of BR signaling in Arabidopsis thaliana has grown steadily since the BR receptor gene BRI1 was cloned in 1997, thanks to genetics, yeast two-hybrid screens, and biochemical and proteomics approaches (reviewed by Jaillais and Vert, 2016), but the position of each component within the cascade is still under refinement. In their article, Kim et al. (2019) assign a role in BR signaling to the U-box E3 ubiquitin ligase PUB40.
Let me introduce the acronyms of some of the players, as illustrated in the Figure. In the absence of BR, BRI1 and its co-receptor BAK1 are physically kept apart by BKI1, which leaves the kinase BIN2 free to phosphorylate the transcriptional effectors of the pathway, BZR1 and its homolog BES1. Phosphorylated BZR1 and BES1 cannot bind target DNA, are kept out of the nucleus, and are targeted for degradation (Jaillais and Vert, 2016).
The perception of BR at the cell surface initiates a kinase/phosphatase cascade that ends with the inactivation and degradation of BIN2: the BRI1-BAK1 complex phosphorylates itself and the downstream kinases CDG1 and BSK1, in turn activating the phosphatase BSU1. The substrate of BSU1 is none other than BIN2, which undergoes activation by autophosphorylation. Non-phosphorylated BZR1 and BES1 can then accumulate in the nucleus and modulate gene expression (Jaillais and Vert, 2016).
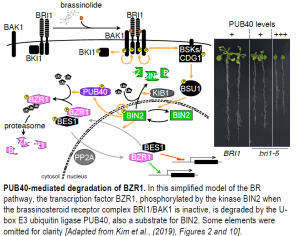 BR signaling borrows elements from the metazoan Wnt/b-catenin pathway (a secreted glycolipoprotein and its associated transcriptional coactivator): both involve a BIN2-type kinase and degradation of the transcriptional effector when the receptor is inactive. b-catenin is composed of Armadillo repeats; almost half of Arabidopsis proteins with Armadillo repeats are E3 ubiquitin ligases. Of a selected subset of these E3 ligases, only PUB40 (and its homologs PUB38, 39, and 41) interacted with BZR1. PUB40, a cytosolic protein, showed a marked preference for phosphorylated BZR1 upon activation of BR signaling and mediated its degradation via the proteasome in roots, but not in leaves. Conversely, the dominant bzr1-1D mutant, which carries a point mutation in the protein’s PEST domain, decreased the interaction between BZR1 and PUB40, resulting in higher BZR1 levels.
BR signaling borrows elements from the metazoan Wnt/b-catenin pathway (a secreted glycolipoprotein and its associated transcriptional coactivator): both involve a BIN2-type kinase and degradation of the transcriptional effector when the receptor is inactive. b-catenin is composed of Armadillo repeats; almost half of Arabidopsis proteins with Armadillo repeats are E3 ubiquitin ligases. Of a selected subset of these E3 ligases, only PUB40 (and its homologs PUB38, 39, and 41) interacted with BZR1. PUB40, a cytosolic protein, showed a marked preference for phosphorylated BZR1 upon activation of BR signaling and mediated its degradation via the proteasome in roots, but not in leaves. Conversely, the dominant bzr1-1D mutant, which carries a point mutation in the protein’s PEST domain, decreased the interaction between BZR1 and PUB40, resulting in higher BZR1 levels.
The authors then confirmed the role of PUB40 as a negative regulator of BR signaling genetically: overexpression of PUB40 in a weak bri1 mutant increased the severity of BR-related phenotypes, while a pub39 pub40 pub41 triple mutant behaved like BZR1 overexpressing lines in terms of root elongation and gene expression profile (see Figure).
Is PUB40 activity controlled by BR? Yes, via BIN2-mediated phosphorylation, which strengthens PUB40’s binding to BZR1, but also marks the E3 ligase for degradation by the proteasome. The function of PUB40 is clearly restricted to roots; another set of players likely function in leaves: perhaps the blue light photoreceptor CRY1 (He et al., 2019)?
References:
He G, Liu J, Dong H, and Sun J (2019). The Blue Light Receptor CRY1 Interacts with BZR1 and BIN2 to Modulate the Phosphorylation and Nuclear Function of BZR1 in Repressing BR Signaling in Arabidopsis. Mol. Plant. doi: https://doi.org/10.1016/j.molp.2019.02.001
Jaillais Y and Vert G (2016). Brassinosteroid signaling and BRI1 dynamics went underground. Curr Opin Plant Biol. 33: 92–100. doi:10.1016/j.pbi.2016.06.014.
Kim E-J, Lee S-H, Park C-H, Kim S-H, Hsu C-C, Xu S, Wang Z-Y, Kim S-K, and Kim T-W (2019). Plant U-Box 40 Mediates Degradation of the Brassinosteroid-Responsive Transcription Factor BZR1 in Arabidopsis Roots. The Plant Cell. Published February 2019. DOI: https://doi.org/10.1105/tpc.18.00941
Levin ER and Hammes SR (2016). Nuclear receptors outside the nucleus: extranuclear signalling by steroid receptors. Nat Rev Mol Cell Biol 17: 783-797. doi: 10.1038/nrm.2016.122.



