Identification of putative substrates of SEC2, a chloroplast inner envelope translocase
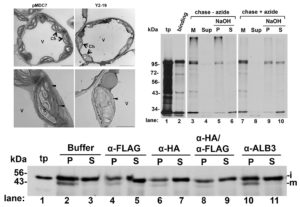 Chloroplasts have evolved from photosynthetic bacteria, and genes necessary for chloroplast function have moved from the chloroplast to the nuclear genome of the host eukaryotic cell. Proteins encoded by these genes are made in the cytosol and imported into the chloroplast using protein translocases. Chloroplasts contain the characterized SEC1 and the recently-discovered SEC2 translocase systems. The SEC2 translocase consists of the SECA2, SCY2, and SECE2 proteins. Since SEC2 was recently discovered, its protein substrates are unknown. Inducible RNAi targeting SCY2 revealed decreased levels of many candidate proteins upon silencing. To identify substrates for SEC2, Li, et al. developed an in vitro method to test protein integration into isolated chloroplast membranes. Using this assay, FTSH12 integration in the assay was inhibited by the SECA inhibitor sodium azide, which identified FTSH12 as a SEC2 substrate. TIC40 was identified as a second substrate because antibodies against epitope-tagged SCY2 and SECE2 inhibited TIC40 integration. (Review by Daniel Czerny) Plant Physiol.
Chloroplasts have evolved from photosynthetic bacteria, and genes necessary for chloroplast function have moved from the chloroplast to the nuclear genome of the host eukaryotic cell. Proteins encoded by these genes are made in the cytosol and imported into the chloroplast using protein translocases. Chloroplasts contain the characterized SEC1 and the recently-discovered SEC2 translocase systems. The SEC2 translocase consists of the SECA2, SCY2, and SECE2 proteins. Since SEC2 was recently discovered, its protein substrates are unknown. Inducible RNAi targeting SCY2 revealed decreased levels of many candidate proteins upon silencing. To identify substrates for SEC2, Li, et al. developed an in vitro method to test protein integration into isolated chloroplast membranes. Using this assay, FTSH12 integration in the assay was inhibited by the SECA inhibitor sodium azide, which identified FTSH12 as a SEC2 substrate. TIC40 was identified as a second substrate because antibodies against epitope-tagged SCY2 and SECE2 inhibited TIC40 integration. (Review by Daniel Czerny) Plant Physiol.






Leave a Reply
Want to join the discussion?Feel free to contribute!